Mouse and human embryonic genome activation initiate at the one-cell stage
- PMID: 40809689
- PMCID: PMC12343560
- DOI: 10.3389/fcell.2025.1594995
Mouse and human embryonic genome activation initiate at the one-cell stage
Abstract
At the moment of their union, fertilizing gametes (sperm and oocyte) are transcriptionally silent: gene expression has to be initiated within the resulting embryo, a process termed embryonic genome activation, EGA. Until recently, EGA was believed to occur at the two-cell stage (mouse) or four-to-eight-cell stage (human), but new evidence from single-cell RNA-sequencing (scRNAseq) suggests that it initiates at the one-cell stage in both species. Precise time-course scRNA-seq of mouse one-cell embryos revealed an EGA program referred to as immediate EGA, iEGA: iEGA occurred from within 4 h of fertilization, mainly from the maternal genome, with paternal genomic transcription from ∼10 h. Significant low-magnitude upregulation similarly occurred in healthy human one-cell embryos. In both species, new transcripts were canonically spliced, and expression predicted embryonic processes and regulatory transcription factors (TFs) associated with cancer, including MYC/c-Myc. Blocking their activities in mouse one-cell embryos induced acute developmental arrest and disrupted iEGA. Inhibiting c-Myc induced upregulation of hundreds of genes, implying that they are normatively repressed, a phenomenon we term embryonic genome repression, EGR. iEGA is downregulated coincidentally with a subsequent, higher-amplitude wave of gene expression (referred to as 'major EGA' or 'major ZGA') in two-cell (mouse) or 4-8-cell (human) embryos. We suggest that iEGA is continuous with gene expression previously termed 'minor EGA' (or 'minor ZGA') and that the regulation of iEGA and major EGA are distinctive. The pattern of gene upregulation in iEGA illuminates processes involved at the onset of development, with implications for epigenetic inheritance, stem cell-derived embryos and cancer.
Keywords: embryonic genome activation (EGA); embryonic genome repression (EGR); fertilization; immediate EGA; one-cell embryo; single-cell RNA-sequencing; transcription; zygotic genome activation (ZGA).
Copyright © 2025 Asami and Perry.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures
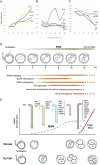
Similar articles
-
Prescription of Controlled Substances: Benefits and Risks.2025 Jul 6. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. 2025 Jul 6. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. PMID: 30726003 Free Books & Documents.
-
A program of successive gene expression in mouse one-cell embryos.Cell Rep. 2023 Feb 28;42(2):112023. doi: 10.1016/j.celrep.2023.112023. Epub 2023 Jan 31. Cell Rep. 2023. PMID: 36729835
-
Maternal transcription factor OTX2 directly induces SETD1A and promotes embryonic genome activation in human pre-implantation embryos.Sci China Life Sci. 2025 Jul;68(7):2058-2072. doi: 10.1007/s11427-024-2875-3. Epub 2025 Apr 22. Sci China Life Sci. 2025. PMID: 40285911
-
The "life code": A theory that unifies the human life cycle and the origin of human tumors.Semin Cancer Biol. 2020 Feb;60:380-397. doi: 10.1016/j.semcancer.2019.09.005. Epub 2019 Sep 12. Semin Cancer Biol. 2020. PMID: 31521747 Review.
-
The Black Book of Psychotropic Dosing and Monitoring.Psychopharmacol Bull. 2024 Jul 8;54(3):8-59. Psychopharmacol Bull. 2024. PMID: 38993656 Free PMC article. Review.
References
LinkOut - more resources
Full Text Sources

