An A. thaliana mutant lacking all nine ATG8 isoforms provides genetic evidence for functional specialization of ATG8 in plants
- PMID: 40814261
- PMCID: PMC12450472
- DOI: 10.1242/jcs.263803
An A. thaliana mutant lacking all nine ATG8 isoforms provides genetic evidence for functional specialization of ATG8 in plants
Abstract
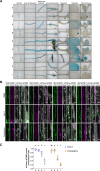
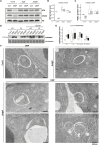
Autophagy sustains cellular health by recycling damaged or excess components through autophagosomes. Autophagy is mediated by conserved ATG proteins, among which the ubiquitin-like ATG8 proteins play a central role by linking cargo to the growing autophagosomes. Unlike most ATG proteins, the ATG8 gene family is significantly expanded in vascular plants, but its functional specialization remains poorly understood. Using transcriptional and translational reporters in Arabidopsis thaliana, we revealed that ATG8 isoforms are differentially expressed across tissues and form distinct autophagosomes. To explore ATG8 specialization, we generated the nonuple Δatg8 mutant, lacking all nine ATG8 isoforms. The mutant displayed hypersensitivity to carbon and nitrogen starvation, coupled with defects in bulk and selective autophagy, as shown by biochemical and ultrastructural analyses. Complementation experiments demonstrated that ATG8A could rescue both carbon and nitrogen starvation phenotypes, whereas ATG8H could only complement carbon starvation. Proximity labeling proteomics further identified isoform-specific interactors under nitrogen starvation, underscoring their functional divergence. These findings provide genetic evidence for functional specialization of ATG8 isoforms in plants and lay the foundation for investigating their roles in diverse cell types and stress conditions.
Keywords: Arabidopsis; Atg8; Autophagy; Selective autophagy.
© 2025. Published by The Company of Biologists.
Conflict of interest statement
Competing interests The authors declare no competing or financial interests.
Figures


References
MeSH terms
Substances
Grants and funding
- P32355/Austrian Science Fund
- P34944/Austrian Science Fund
- SFB F79/Austrian Science Fund
- DOC 111/Austrian Science Fund
- LS17-47/Vienna Science and Technology Fund
- LS21-09/Vienna Science and Technology Fund
- 101043370/HORIZON EUROPE European Research Council
- Heisenberg Professorship/Deutsche Forschungsgemeinschaft
- 847548/HORIZON EUROPE Marie Sklodowska-Curie Actions
- LS17-047/Vienna Science and Technology Fund
- LS21-009/Vienna Science and Technology Fund
- 101043370/ERC_/European Research Council/International
- Deutsche Forschungsgemeinschaft
- Austrian Academy of Sciences
LinkOut - more resources
Full Text Sources

