Global trends in machine learning applications for single-cell transcriptomics research
- PMID: 40819059
- PMCID: PMC12357469
- DOI: 10.1186/s41065-025-00528-y
Global trends in machine learning applications for single-cell transcriptomics research
Abstract
Background: Single-cell RNA sequencing (scRNA-seq) has revolutionized cellular heterogeneity analysis by decoding gene expression profiles at individual cell level, while machine learning (ML) has emerged as core computational tool for clustering analysis, dimensionality reduction modeling and developmental trajectory inference in single-cell transcriptomics(SCT). Although 3,307 papers have been published in past two decades, there remains lack of bibliometric review comprehensively addressing methodological evolution, technical challenges and clinical translation pathways. This study aims to fill research gap through bibliometric and visual analysis, revealing technological evolution trends and future development directions.
Methods: Using 3,307 publications from Web of Science Core Collection(WOSCC), we conducted bibliometric and visualization analysis through CiteSpace and VOSviewer to systematically review research trends, national/institutional contributions, keyword co-occurrence networks and co-citation relationships. Data screening strictly limited to English articles and reviews, excluding irrelevant document types, focusing on core application scenarios of ML in SCT.
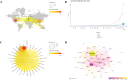
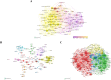
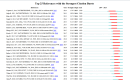
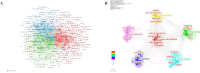
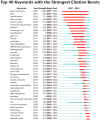
Results: China and United States dominated research output (combined 65%), with China leading in publication volume (54.8%) while US demonstrating academic influence through H-index 84 and 37,135 total citations. Research hotspots concentrated on random forest (RF) and deep learning models, showing transition from algorithm development to clinical applications (e.g., tumor immune microenvironment analysis). Chinese Academy of Sciences and Harvard University emerged as core collaboration hubs, with international cooperation network primarily featuring US-China collaboration. Keyword clustering revealed four themes: gene expression, immunotherapy, bioinformatics, and inflammation-related research. Technical bottlenecks included data heterogeneity, insufficient model interpretability and weak cross-dataset generalization capability.
Conclusion: ML-scRNA-seq integration has advanced cellular heterogeneity analysis and precision medicine development. Future directions should optimize deep learning architectures, enhance model generalization capabilities, and promote technical translation through multi-omics and clinical data integration. Interdisciplinary collaboration represents key to overcoming current limitations (e.g., data standardization, algorithm interpretability), ultimately realizing deep integration between single-cell technologies and precision medicine.
Keywords: Bibliometric analysis; Deep learning; Machine learning; Random forest; Single-cell transcriptomics.
© 2025. The Author(s).
Conflict of interest statement
Declarations. Ethics approval and consent to participate: Not applicable. Consent for publication: Not applicable. Competing interests: The authors declare no competing interests.
Figures


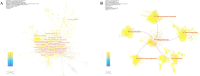

Similar articles
-
Data-driven trends in critical care informatics: a bibliometric analysis of global collaborations using the MIMIC database (2004-2024).Comput Biol Med. 2025 Sep;195:110670. doi: 10.1016/j.compbiomed.2025.110670. Epub 2025 Jun 27. Comput Biol Med. 2025. PMID: 40580617
-
Driving innovations in cancer research through spatial metabolomics: a bibliometric review of trends and hotspot.Front Immunol. 2025 Jun 10;16:1589943. doi: 10.3389/fimmu.2025.1589943. eCollection 2025. Front Immunol. 2025. PMID: 40557160 Free PMC article.
-
Research status, hotspots and perspectives of artificial intelligence applied to pain management: a bibliometric and visual analysis.Updates Surg. 2025 Jun 28. doi: 10.1007/s13304-025-02296-w. Online ahead of print. Updates Surg. 2025. PMID: 40580377
-
A bibliometric analysis of research trends in mesenchymal stem cell therapy for neonatal bronchopulmonary dysplasia: 2004-2024.Front Pediatr. 2025 Jun 3;13:1558301. doi: 10.3389/fped.2025.1558301. eCollection 2025. Front Pediatr. 2025. PMID: 40530182 Free PMC article. Review.
-
Global research on Chinese martial arts (1974-2025): A bibliometric and visualization-based analysis using Web of Science.Medicine (Baltimore). 2025 Aug 8;104(32):e43769. doi: 10.1097/MD.0000000000043769. Medicine (Baltimore). 2025. PMID: 40797508 Free PMC article.
References
-
- Wani SA, Khan SA. and S.J.A.o.C.M.i.E. Quadri, Application of deep learning for single cell multi-omics: a state-of-the-art review. 2025: pp. 1–43.
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

