Multi-omics analysis of Taiwanofungus gaoligongensis: effects of different cultivation methods on secondary metabolites
- PMID: 40822405
- PMCID: PMC12355660
- DOI: 10.3389/fmicb.2025.1620693
Multi-omics analysis of Taiwanofungus gaoligongensis: effects of different cultivation methods on secondary metabolites
Abstract
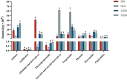
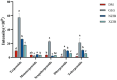
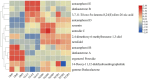
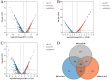
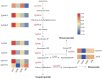
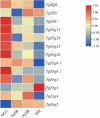
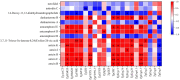
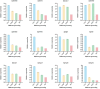
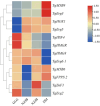
A multi-omics strategy was utilized in this study to investigate the effects of various cultivation methods-including the fruiting bodies cultivation on Cinnamomum kanehirae wood logs (GLG), the mycelia cultivation on C. kanehirae substrate fungal cultivation bags (NZJB), Cinnamomum camphora substrate fungal cultivation bags (XZJB) and rice medium (DM)-on Secondary Metabolites in Taiwanofungus gaoligongensis. NZJB and XZJB significantly enhanced terpenoids production in the mycelium, with triterpenoid contents in NZJB and XZJB being sevenfold and 3.9-fold higher, respectively, than those in DM. Antcins were notably increased in fungal cultivation bag cultures: antcin C reached the highest level in XZJB (9.72-fold higher than in DM), antcin I peaked in NZJB (12.83-fold higher than in DM), and antrodin C also reached its maximum in NZJB. Additionally, the antrodin C content in NZJB was 3.2-fold higher than in GLG and 4.08-fold higher than in DM. In addition, the levels of steroids, phenolic compounds, and flavonoids were also significantly increased in NZJB and XZJB. Transcriptome analysis revealed significant differences in the expression of genes involved in the biosynthesis of antcins and antrodin C across the different cultivation methods. In particular, the expression of TgHMGR was markedly higher in NZJB than in XZJB and DM, correlating with the elevated terpenoids and triterpenoids levels, suggesting that TgHMGR may act as a key rate-limiting enzyme in the terpenoid biosynthesis pathway of T. gaoligongensis. The expression levels of terpenoid biosynthesis-related genes were significantly elevated in GLG compared to mycelium, consistent with the higher abundance of terpenoid metabolites. Co-expression analysis of transcription factors (TFs) and promoter binding site predictions indicated that the expression of TgHMGR and TgFPPS 2 may be regulated by TgHSF4 and TgMYB6, respectively. Meanwhile, the expression of TgErg2, TgErg3, TgErg5, and TgErg6 1 may be regulated by TgZnF1, TgMYB9, TgHOX1, and TgHMG8. This study compared the metabolite profiles and gene expression patterns of the fruiting bodies of T. gaoligongensis with those of three types of cultivated mycelia. The results provide new insights into the transcriptional regulation of key bioactive compound biosynthesis in T. gaoligongensis and suggest potential strategies to enhance the production of active compounds in mycelia through artificial cultivation, thereby improving its medicinal value and production efficiency.
Keywords: Taiwanofungus gaoligongensis; reference-based transcriptomic analysis; secondary metabolites; transcription factor; transcriptional regulation; untargeted metabolomic analysis.
Copyright © 2025 He, Yuan, Xiao, Hu, Wang, Zhao, Li, Peng, Zhang and Zheng.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures


Similar articles
-
Prescription of Controlled Substances: Benefits and Risks.2025 Jul 6. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. 2025 Jul 6. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. PMID: 30726003 Free Books & Documents.
-
Systemic treatments for metastatic cutaneous melanoma.Cochrane Database Syst Rev. 2018 Feb 6;2(2):CD011123. doi: 10.1002/14651858.CD011123.pub2. Cochrane Database Syst Rev. 2018. PMID: 29405038 Free PMC article.
-
Systematic review on urine albumin testing for early detection of diabetic complications.Health Technol Assess. 2005 Aug;9(30):iii-vi, xiii-163. doi: 10.3310/hta9300. Health Technol Assess. 2005. PMID: 16095545
-
Genomic Features of Taiwanofungus gaoligongensis and the Transcriptional Regulation of Secondary Metabolite Biosynthesis.J Fungi (Basel). 2024 Nov 27;10(12):826. doi: 10.3390/jof10120826. J Fungi (Basel). 2024. PMID: 39728323 Free PMC article.
-
Sexual Harassment and Prevention Training.2024 Mar 29. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. 2024 Mar 29. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. PMID: 36508513 Free Books & Documents.
References
-
- Achudhan D., Liu S., Lin Y., Huang C., Tsai C., Ko C., et al. (2021). Antcin K inhibits TNF-alpha, IL-1beta and IL-8 expression in synovial fibroblasts and ameliorates cartilage degradation: Implications for the treatment of rheumatoid arthritis. Front. Immunol. 12:790925. 10.3389/fimmu.2021.790925 - DOI - PMC - PubMed
-
- Adiguzel O., Li J., Lin X., Shao E., Lin Z., Zhi-Gang T., et al. (2016). Molecular cloning and expression analysis of mevalonate pyrophosphate decarboxylase in Antrodia cinnamomea. MATEC Web Confer. 64:03001. 10.1051/matecconf/20166403001 - DOI
LinkOut - more resources
Full Text Sources

