Multiple golgins are required to support extracellular matrix secretion, modification, and assembly
- PMID: 40824304
- PMCID: PMC12360289
- DOI: 10.1083/jcb.202411167
Multiple golgins are required to support extracellular matrix secretion, modification, and assembly
Abstract
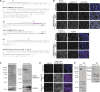
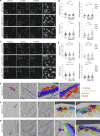
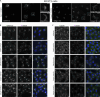
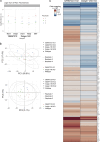
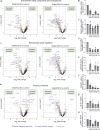
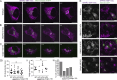
The secretion of extracellular matrix (ECM) proteins is vital to the maintenance of tissue health. One major control point of this process is the Golgi apparatus, whose dysfunction causes numerous connective tissue disorders. We therefore sought to investigate the role of two Golgi organizing proteins, GMAP210 and Golgin-160, in ECM secretion. CRISPR knockout of either protein had distinct impacts on Golgi organization, with Golgin-160 knockout causing Golgi fragmentation and vesicle accumulation, and GMAP210 loss leading to cisternal fragmentation, dilation, and the accumulation of tubulovesicular structures. Both golgins were required for fibrillar collagen organization and glycosaminoglycan synthesis suggesting nonredundant functions in these processes. Furthermore, proteomics analysis revealed both shared and golgin-specific changes in the secretion of ECM proteins. We therefore propose that golgins are collectively required to create the correct physical-chemical space to support efficient ECM protein secretion, modification, and assembly. This is the first time that Golgin-160 has been shown to be required for ECM secretion.
© 2025 Thompson et al.
Conflict of interest statement
Disclosures: The authors declare no competing interests exist.
Figures







Similar articles
-
Prediction, screening and characterization of novel bioactive tetrapeptide matrikines for skin rejuvenation.Br J Dermatol. 2024 Jun 20;191(1):92-106. doi: 10.1093/bjd/ljae061. Br J Dermatol. 2024. PMID: 38375775
-
Prescription of Controlled Substances: Benefits and Risks.2025 Jul 6. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. 2025 Jul 6. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. PMID: 30726003 Free Books & Documents.
-
Golgin Protein MoCoy1 Mediates Retrograde Trafficking From the Golgi to the ER, Regulating Fungal Development and Pathogenicity in Magnaporthe oryzae.Mol Plant Pathol. 2025 Jul;26(7):e70130. doi: 10.1111/mpp.70130. Mol Plant Pathol. 2025. PMID: 40700342 Free PMC article.
-
Golgi's Role in the Development of Possible New Therapies in Cancer.Cells. 2023 May 29;12(11):1499. doi: 10.3390/cells12111499. Cells. 2023. PMID: 37296620 Free PMC article. Review.
-
Comparison of Natural Polysaccharides and Synthetic Polymers in Facilitating Macromolecular Crowding to Create Customized Extracellular Matrix-Rich Supramolecular Assemblies from Fibroblasts.ACS Appl Bio Mater. 2025 Aug 18;8(8):6701-6721. doi: 10.1021/acsabm.5c01003. Epub 2025 Aug 5. ACS Appl Bio Mater. 2025. PMID: 40763242 Review.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

