Mechanistic computational modeling of sFLT1 secretion dynamics
- PMID: 40825055
- PMCID: PMC12370208
- DOI: 10.1371/journal.pcbi.1013324
Mechanistic computational modeling of sFLT1 secretion dynamics
Abstract
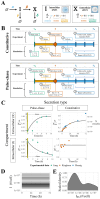
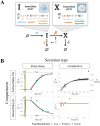
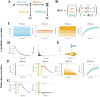
Constitutively secreted by endothelial cells, soluble FLT1 (sFLT1 or sVEGFR1) binds and sequesters extracellular vascular endothelial growth factors (VEGF), thereby reducing VEGF binding to VEGF receptor tyrosine kinases and their downstream signaling. In doing so, sFLT1 plays an important role in vascular development and in the patterning of new blood vessels in angiogenesis. Here, we develop multiple mechanistic models of sFLT1 secretion and identify a minimal mechanistic model that recapitulates key qualitative and quantitative features of temporal experimental datasets of sFLT1 secretion from multiple studies. We show that the experimental data on sFLT1 secretion is best represented by a delay differential equation (DDE) system including a maturation term, reflecting the time required between synthesis and secretion. Using optimization to identify appropriate values for the key mechanistic parameters in the model, we show that two model parameters (extracellular degradation rate constant and maturation time) are very strongly constrained by the experimental data, and that the remaining parameters are related by two strongly constrained constants. Thus, only one degree of freedom remains, and measurements of the intracellular levels of sFLT1 would fix the remaining parameters. Comparison between simulation predictions and additional experimental data of the outcomes of chemical inhibitors and genetic perturbations suggest that intermediate values of the secretion rate constant best match the simulation with experiments, which would completely constrain the model. However, some of the inhibitors tested produce results that cannot be reproduced by the model simulations, suggesting that additional mechanisms not included here are required to explain those inhibitors. Overall, the model reproduces most available experimental data and suggests targets for further quantitative investigation of the sFLT1 system.
Copyright: © 2025 Gill et al. This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures





Update of
-
Mechanistic computational modeling of sFLT1 secretion dynamics.bioRxiv [Preprint]. 2025 Feb 18:2025.02.12.637983. doi: 10.1101/2025.02.12.637983. bioRxiv. 2025. Update in: PLoS Comput Biol. 2025 Aug 18;21(8):e1013324. doi: 10.1371/journal.pcbi.1013324. PMID: 40027776 Free PMC article. Updated. Preprint.
Similar articles
-
Mechanistic computational modeling of sFLT1 secretion dynamics.bioRxiv [Preprint]. 2025 Feb 18:2025.02.12.637983. doi: 10.1101/2025.02.12.637983. bioRxiv. 2025. Update in: PLoS Comput Biol. 2025 Aug 18;21(8):e1013324. doi: 10.1371/journal.pcbi.1013324. PMID: 40027776 Free PMC article. Updated. Preprint.
-
Impact of ligand binding on VEGFR1, VEGFR2, and NRP1 localization in human endothelial cells.PLoS Comput Biol. 2025 Jul 16;21(7):e1013254. doi: 10.1371/journal.pcbi.1013254. eCollection 2025 Jul. PLoS Comput Biol. 2025. PMID: 40668864 Free PMC article.
-
Prescription of Controlled Substances: Benefits and Risks.2025 Jul 6. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. 2025 Jul 6. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. PMID: 30726003 Free Books & Documents.
-
Anti-vascular endothelial growth factor biosimilars for neovascular age-related macular degeneration.Cochrane Database Syst Rev. 2024 Jun 3;6(6):CD015804. doi: 10.1002/14651858.CD015804.pub2. Cochrane Database Syst Rev. 2024. PMID: 38829176 Free PMC article.
-
The Black Book of Psychotropic Dosing and Monitoring.Psychopharmacol Bull. 2024 Jul 8;54(3):8-59. Psychopharmacol Bull. 2024. PMID: 38993656 Free PMC article. Review.
References
-
- Uhlén M, et al. The human secretome. Sci Signal. 2019;12:eaaz0274. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

