Tissue-Specific Immunosuppressive and Proliferating Macrophages Fuel Early Metastatic Progression of Human Colorectal Cancer to the Liver
- PMID: 40828155
- PMCID: PMC12580778
- DOI: 10.1158/2326-6066.CIR-25-0031
Tissue-Specific Immunosuppressive and Proliferating Macrophages Fuel Early Metastatic Progression of Human Colorectal Cancer to the Liver
Abstract
Early synchronous colorectal liver metastasis (CRLM) represents a clinical condition characterized by the simultaneous presence of primary colorectal cancer and metastatic liver lesions. In this study, we characterized the tissue-specific transcriptomes, phenotypes, and functional relevance of tumor-associated macrophages (TAM) within the tumor microenvironment (TME) of colorectal cancer and CRLM specimens from patients who underwent simultaneous surgical removal of these malignancies. The high-throughput single-cell transcriptional analysis revealed an inverse ratio of inflammatory and immunoregulatory TAMs in the colorectal cancer and CRLM TMEs, along with heterogeneity in both tumoral tissues. Furthermore, we found that inflammatory TAMs in colorectal cancer expressed inhibitory ligands that might support immune escape, thus favoring liver metastatic progression. In contrast, CRLM lesions possessed a highly immunosuppressive milieu characterized by large proliferative CTLA4+ immunoregulatory TAMs and the presence of IL7R+ cytotoxic TAMs. Higher frequencies of these specific TAM subsets in CRLM were associated with shorter disease-free survival and worse patient prognosis. The identification and characterization of immunoregulatory TAMs preferentially enriched in CRLM is key for the development of novel immunotherapeutic strategies aimed at boosting anticancer immune responses within the TME.
©2025 The Authors; Published by the American Association for Cancer Research.
Conflict of interest statement
F. Milana reports grants from the Ministry of Health during the conduct of the study. No disclosures were reported by the other authors.
Figures




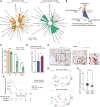
References
-
- Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 2021;71:209–49. - PubMed
-
- Cucchetti A, Ferrero A, Cescon M, Donadon M, Russolillo N, Ercolani G, et al. Cure model survival analysis after hepatic resection for colorectal liver metastases. Ann Surg Oncol 2015;22:1908–14. - PubMed
-
- Siriwardena AK, Serrablo A, Fretland ÅA, Wigmore SJ, Ramia-Angel JM, Malik HZ, et al. Multisocietal European consensus on the terminology, diagnosis, and management of patients with synchronous colorectal cancer and liver metastases: an E-AHPBA consensus in partnership with ESSO, ESCP, ESGAR, and CIRSE. Br J Surg 2023;110:1161–70. - PMC - PubMed

