Ferroptosis-related hub genes and immune cell dynamics as diagnostic biomarkers in age-related macular degeneration
- PMID: 40830492
- PMCID: PMC12366384
- DOI: 10.1186/s40001-025-03044-x
Ferroptosis-related hub genes and immune cell dynamics as diagnostic biomarkers in age-related macular degeneration
Abstract
Background: Age-related Macular Degeneration (AMD) is widely acknowledged as a principal cause of vision loss in the elderly. Currently, the therapeutic interventions available in clinical practice fail to achieve satisfactory outcomes. Therefore, it is imperative that we approach the progress of AMD from novel perspectives in order to explore new therapeutic strategies.
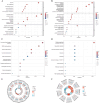
Method: We obtained transcriptomic data from the macular and the peripheral retina from patients with AMD and a control group from the Gene Expression Omnibus (GEO) database. Through Gene Ontology (GO) analysis and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis, we identified differentially expressed genes (DEGs) that were significantly enriched in functions associated with ferroptosis. Subsequent application of machine learning techniques enabled the identification of key hub genes, whose diagnostic potential was further validated. Additionally, the expression of these hub genes was corroborated in both animal and cellular models. Finally, we performed a functional enrichment analysis of these hub genes.
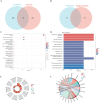
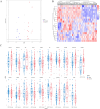
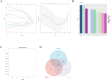
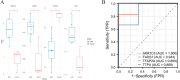
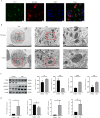
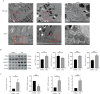
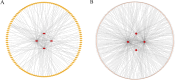
Results: In the macula of patients with AMD, 452 DEGs were identified, while in the peripheral retina, 222 DEGs were discovered. Within the macula, 19 genes were associated with ferroptosis, compared to 3 in the peripheral retina. Consequently, the macular was selected as the primary focus of the study. Subsequent screening of these 19 genes using LASSO regression, Support Vector Machine (SVM), and Random Forest algorithms identified four hub genes: FADS1, TFAP2A, AKR1C3, and TTPA. Consequently, we utilized cigarette smoke extract (CSE) to either stimulate retinal pigment epithelial (RPE) cells in vitro or administer it via intravitreal injection, thereby establishing in vitro and in vivo models of AMD. Results from RT-PCR and Western blot analyses revealed an upregulation of FADS1, AKR1C3, and TTPA, while TFAP2A exhibited decreased expression. Finally, we investigated the infiltration of immune cells within the macular and performed a functional enrichment analysis of the hub genes.
Conclusion: We identified four key ferroptosis-related genes (FRGs)-FADS1, AKR1C3, TFAP2A, and TTPA-that possess diagnostic relevance for AMD and correlate with immune cell infiltration. Moreover, significant changes in both mRNA and protein expression levels of these genes have been observed in in vitro experiments and mice models.
Keywords: Age-related macular degeneration; Ferroptosis-related genes; Immune cells; Machine learning.
© 2025. The Author(s).
Conflict of interest statement
Declarations. Ethics approval and consent to participate: This study received ethical approval from the Ethics Committee of The First Affiliated Hospital of Chongqing Medical University (Chongqing, China), with the approval number IACUC-CQMU-2023-0355. Competing interests: The authors declare no competing interests.
Figures




Similar articles
-
Identification of osteoporosis ferroptosis-related markers and potential therapeutic compounds based on bioinformatics methods and molecular docking technology.BMC Med Genomics. 2024 Apr 22;17(1):99. doi: 10.1186/s12920-024-01872-0. BMC Med Genomics. 2024. PMID: 38650009 Free PMC article.
-
Machine learning based screening of biomarkers associated with cell death and immunosuppression of multiple life stages sepsis populations.Sci Rep. 2025 Aug 19;15(1):30302. doi: 10.1038/s41598-025-14600-0. Sci Rep. 2025. PMID: 40830558 Free PMC article.
-
Artificial intelligence for diagnosing exudative age-related macular degeneration.Cochrane Database Syst Rev. 2024 Oct 17;10(10):CD015522. doi: 10.1002/14651858.CD015522.pub2. Cochrane Database Syst Rev. 2024. PMID: 39417312
-
Deciphering Shared Gene Signatures and Immune Infiltration Characteristics Between Gestational Diabetes Mellitus and Preeclampsia by Integrated Bioinformatics Analysis and Machine Learning.Reprod Sci. 2025 Jun;32(6):1886-1904. doi: 10.1007/s43032-025-01847-1. Epub 2025 May 15. Reprod Sci. 2025. PMID: 40374866
-
Blue-light filtering intraocular lenses (IOLs) for protecting macular health.Cochrane Database Syst Rev. 2018 May 22;5(5):CD011977. doi: 10.1002/14651858.CD011977.pub2. Cochrane Database Syst Rev. 2018. PMID: 29786830 Free PMC article.
References
-
- Singh A, Faber C, Falk M, Nissen MH, Hviid TV, Sørensen TL. Altered expression of CD46 and CD59 on leukocytes in neovascular age-related macular degeneration. Am J Ophthalmol. 2012. 10.1016/j.ajo.2012.01.036. - PubMed
-
- Zhang N, Peairs JJ, Yang P, et al. The importance of Bcl-xL in the survival of human RPE cells. Invest Ophthalmol Vis Sci. 2007;48(8):3846–53. 10.1167/iovs.06-1145. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

