Construction of a feature gene and machine prediction model for inflammatory bowel disease based on multichip joint analysis
- PMID: 40830794
- PMCID: PMC12366088
- DOI: 10.1186/s12967-025-06838-z
Construction of a feature gene and machine prediction model for inflammatory bowel disease based on multichip joint analysis
Abstract
Background: Inflammatory bowel disease (IBD) is a chronic nonspecific inflammatory disorder triggered by immune responses and genetic factors. Currently, there is no cure for IBD, and its etiology remains unclear. As a result, early detection and diagnosis of IBD pose significant challenges. Therefore, investigating biomarkers in peripheral blood is highly important, as they can assist doctors in the early identification and management of IBD.
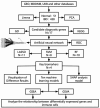
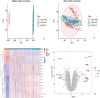
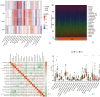
Methods: We used a multichip joint analysis approach to explore the database thoroughly. On the basis of methods such as artificial neural networks (ANNs), machine learning techniques, and the SHAP model, we developed a diagnostic model for IBD. To select genetic features, we utilized three machine learning algorithms, namely, least absolute shrinkage and selection operator (LASSO), support vector machine (SVM), and random forest (RF), to identify differentially expressed genes. Additionally, we conducted an in-depth analysis of the enriched molecular pathways of these differentially expressed genes through Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analyses. Moreover, we used the SHAP model to interpret the results of the machine learning process. Finally, we examined the relationships between the differentially expressed genes and immune cells.
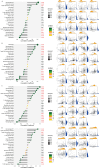
Results: Through machine learning, we identified four crucial biomarkers for IBD, namely, LOC389023, DUOX2, LCN2, and DEFA6. The SHAP model was used to elucidate the contribution of the differentially expressed genes to the diagnostic model. These genes were associated primarily with immune system modulation and microbial alterations. GO and KEGG pathway enrichment analyses indicated that the differentially expressed genes demonstrated associations with molecular pathways such as the antimicrobial and IL-17 signaling pathways. By performing correlation and differential analyses between differentially expressed genes and immune cells, we found that M1 macrophages exhibited stable differential changes in all four differentially expressed genes. M2 macrophages, resting mast cells, neutrophils, and activated memory CD4 T cells all showed significant differences in three of the differentially expressed genes.
Conclusion: We identified differentially expressed genes (LOC389023, DUOX2, LCN2, and DEFA6) with significant immune-related effects in IBD. Our findings suggest that machine learning algorithms outperform ANNs in the diagnosis of IBD. This research provides a theoretical foundation for the clinical diagnosis, targeted therapy, and prognostic evaluation of IBD.
Keywords: Artificial neural network; Diagnostic model; Immune differences; Inflammatory bowel disease; Machine learning.
© 2025. The Author(s).
Conflict of interest statement
Declarations. Ethics approval and consent to participate: According to local legislation and institutional requirements, research on human participants requires ethical review and approval. In compliance with the ethical requirements of Jiangnan University Affiliated Hospital in Wuxi, Jiangsu Province, the ethical approval number for this study is LS2024012. Consent for publication: Not required. Competing interests: The author declares that there are no conflicts of interest.
Figures





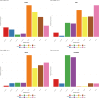
Similar articles
-
Machine learning based screening of biomarkers associated with cell death and immunosuppression of multiple life stages sepsis populations.Sci Rep. 2025 Aug 19;15(1):30302. doi: 10.1038/s41598-025-14600-0. Sci Rep. 2025. PMID: 40830558 Free PMC article.
-
Deciphering Shared Gene Signatures and Immune Infiltration Characteristics Between Gestational Diabetes Mellitus and Preeclampsia by Integrated Bioinformatics Analysis and Machine Learning.Reprod Sci. 2025 Jun;32(6):1886-1904. doi: 10.1007/s43032-025-01847-1. Epub 2025 May 15. Reprod Sci. 2025. PMID: 40374866
-
Prescription of Controlled Substances: Benefits and Risks.2025 Jul 6. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. 2025 Jul 6. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. PMID: 30726003 Free Books & Documents.
-
Patient education interventions for the management of inflammatory bowel disease.Cochrane Database Syst Rev. 2023 May 4;5(5):CD013854. doi: 10.1002/14651858.CD013854.pub2. Cochrane Database Syst Rev. 2023. PMID: 37172140 Free PMC article.
-
Signs and symptoms to determine if a patient presenting in primary care or hospital outpatient settings has COVID-19.Cochrane Database Syst Rev. 2022 May 20;5(5):CD013665. doi: 10.1002/14651858.CD013665.pub3. Cochrane Database Syst Rev. 2022. PMID: 35593186 Free PMC article.
References
-
- Hodson R. Inflammatory bowel disease[J/OL]. Nature. 2016;540(7634):S97–97. 10.1038/540S97a - PubMed
-
- Bisgaard T H, Allin K H Keeferl, et al. Depression and anxiety in inflammatory bowel disease: epidemiology, mechanisms and treatment[J/OL]. Nat Reviews Gastroenterol Hepatol. 2022;19(11):717–26. 10.1038/s41575-022-00634-6 - PubMed
-
- Eftekhar Z, Aghaei M, Saki N. DNA damage repair in megakaryopoiesis: molecular and clinical aspects[J/OL]. Expert Rev Hematol. 2024;17(10):705–12. 10.1080/17474086.2024.2391102 - PubMed
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

