Activation of mitophagy and proteasomal degradation confers resistance to developmental defects in postnatal skeletal muscle
- PMID: 40830874
- PMCID: PMC12362882
- DOI: 10.1186/s12929-025-01153-7
Activation of mitophagy and proteasomal degradation confers resistance to developmental defects in postnatal skeletal muscle
Abstract
Background: Postnatal skeletal muscle development leads to increased muscle mass, strength, and mitochondrial function, but the role of mitochondrial remodeling during this period is unclear. This study investigates mitochondrial remodeling during postnatal muscle development and examines how constitutive autophagy deficiency impacts these processes.
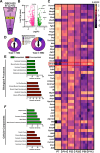
Methods: We initially performed a broad RNA-Seq analysis using a publicly available GEO database of skeletal muscle from postnatal day 7 (P7) to postnatal day 112 (P112) to identify differentially expressed genes. This was followed by investigation of postnatal skeletal muscle development using the mitophagy report mouse line (mt-Kiema mice), as well as conditional skeletal muscle knockout (Atg7f/f:Acta1-Cre) mice.
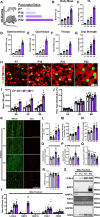
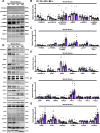
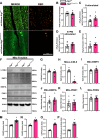
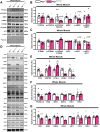
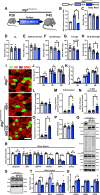
Results: Our study observed rapid growth of body and skeletal muscle mass, along with increased fiber cross-sectional area and grip strength. Mitochondrial maturation was indicated by enhanced maximal respiration, reduced electron leak, and elevated mitophagic flux, as well as increased mitochondrial localization of autophagy and mitophagy proteins. Anabolic signaling was also upregulated, coinciding with increased mitophagy and fusion signaling, and decreased biogenesis signaling. Despite the loss of mitophagic flux in skeletal muscle-specific Atg7 knockout mice, there were no changes in body or skeletal muscle mass; however, hypertrophy was observed in type IIX fibers. This lack of Atg7 and loss of mitophagy was associated with the activation of mitochondrial apoptotic signaling as well as ubiquitin-proteasome signaling, suggesting a shift in degradation mechanisms. Inhibition of the ubiquitin-proteasome system (UPS) in autophagy-deficient skeletal muscle led to significant atrophy, increased reactive oxygen species production, and mitochondrial apoptotic signaling.
Conclusion: These results highlight the role of mitophagy in postnatal skeletal muscle development and suggest that autophagy-deficiency triggers compensatory degradative pathways (i.e., UPS) to prevent mitochondrial apoptotic signaling and thus preserve skeletal muscle integrity in developing mice.
Keywords: Apoptosis; Autophagy; BNIP3; Development; Mitochondria; Mitophagy; Skeletal muscle; UPS.
© 2025. The Author(s).
Conflict of interest statement
Declarations. Ethics approval and consent to participate: All animal procedures were conducted in accordance with the standards set by the Canadian Council on Animal Care (CCAC) and were approved by the Animal Care Committee (ACC) at the University of Waterloo. Consent for publication: Not applicable. Competing interests: The authors declare that they have no competing interests.
Figures

Similar articles
-
Limiting cap-dependent translation increases 20S proteasomal degradation and protects the proteomic integrity in autophagy-deficient skeletal muscle.Autophagy. 2025 Jun;21(6):1212-1227. doi: 10.1080/15548627.2025.2457925. Epub 2025 Feb 6. Autophagy. 2025. PMID: 39878121 Free PMC article.
-
Umbelliferone attenuates diabetic sarcopenia by modulating mitochondrial quality and the ubiquitin-proteasome system.Phytomedicine. 2025 Aug;144:156930. doi: 10.1016/j.phymed.2025.156930. Epub 2025 May 31. Phytomedicine. 2025. PMID: 40483791
-
Impaired Autophagic Flux in Skeletal Muscle of Plectin-Related Epidermolysis Bullosa Simplex With Muscular Dystrophy.J Cachexia Sarcopenia Muscle. 2025 Aug;16(4):e70001. doi: 10.1002/jcsm.70001. J Cachexia Sarcopenia Muscle. 2025. PMID: 40641151 Free PMC article.
-
Physical exercise training interventions for children and young adults during and after treatment for childhood cancer.Cochrane Database Syst Rev. 2016 Mar 31;3(3):CD008796. doi: 10.1002/14651858.CD008796.pub3. Cochrane Database Syst Rev. 2016. PMID: 27030386 Free PMC article.
-
Physical exercise training interventions for children and young adults during and after treatment for childhood cancer.Cochrane Database Syst Rev. 2013 Apr 30;(4):CD008796. doi: 10.1002/14651858.CD008796.pub2. Cochrane Database Syst Rev. 2013. Update in: Cochrane Database Syst Rev. 2016 Mar 31;3:CD008796. doi: 10.1002/14651858.CD008796.pub3. PMID: 23633361 Updated.
References
-
- Gokhin DS, Ward SR, Bremner SN, Lieber RL. Quantitative analysis of neonatal skeletal muscle functional improvement in the mouse. J Exp Biol. 2008;211(Pt 6):837–43. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

