This is a preprint.
A non-catalytic role for RFC in PCNA-mediated processive DNA synthesis
- PMID: 40832337
- PMCID: PMC12363810
- DOI: 10.1101/2025.08.08.669392
A non-catalytic role for RFC in PCNA-mediated processive DNA synthesis
Update in
-
A non-catalytic role for RFC in PCNA-mediated processive DNA synthesis.Cell. 2026 Jan 28:S0092-8674(25)01478-3. doi: 10.1016/j.cell.2025.12.029. Online ahead of print. Cell. 2026. PMID: 41610851 Free PMC article.
Abstract
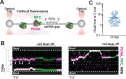
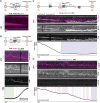
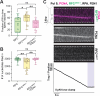
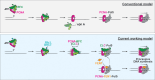
The ring-shaped sliding clamp PCNA enables DNA polymerases to perform processive DNA synthesis during replication and repair. The loading of PCNA onto DNA is catalyzed by the ATPase clamp loader RFC. Using a single-molecule platform to visualize the dynamic interplay between PCNA and RFC on DNA, we unexpectedly discovered that RFC continues to associate with PCNA after loading, contrary to the conventional view. Functionally, this clamp-loader/clamp complex is required for processive DNA synthesis by polymerase δ (Polδ), as the PCNA-Polδ assembly is inherently unstable. This architectural role of RFC is dependent on the BRCT domain of Rfc1, and mutation of its DNA-binding residues causes sensitivity to DNA damage in vivo. We further showed the FEN1 flap endonuclease can also stabilize the PCNA-Polδ interaction and mediate robust synthesis. Overall, our work revealed that, beyond their canonical enzymatic functions, PCNA-binding proteins harbor non-catalytic functions essential for DNA replication and genome maintenance.
Keywords: Clamp loader; DNA damage; DNA replication; FEN1; Fill-in synthesis; Genome maintenance; Okazaki fragment; PCNA; Polδ; RFC; Sliding clamp.
Conflict of interest statement
DECLARATION OF INTERESTS The authors declare no competing interests.
Figures



References
-
- Stillman B., Smart machines at the DNA replication fork. Cell, 1994. 78(5): p. 725–728. - PubMed
-
- Benkovic S.J., Valentine A.M., and Salinas F., Replisome-Mediated DNA Replication. Annual Review of Biochemistry, 2001. 70(Volume 70, 2001): p. 181–208. - PubMed
-
- Johnson A. and O’Donnell M., Cellular DNA replicases: components and dynamics at the replication fork. Annu Rev Biochem, 2005. 74: p. 283–315. - PubMed
-
- Krishna T.S.R., et al. , Crystal structure of the eukaryotic DNA polymerase processivity factor PCNA. Cell, 1994. 79(7): p. 1233–1243. - PubMed
-
- Gulbis J.M., et al. , Structure of the C-terminal region of p21(WAF1/CIP1) complexed with human PCNA. Cell, 1996. 87(2): p. 297–306. - PubMed
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous
