In-silico screening of small compounds against Lassa fever haemorrhagic virus nucleoprotein
- PMID: 40835743
- PMCID: PMC12368151
- DOI: 10.1038/s41598-025-89989-9
In-silico screening of small compounds against Lassa fever haemorrhagic virus nucleoprotein
Abstract
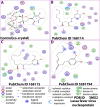
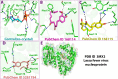
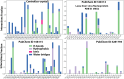
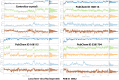
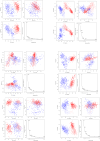
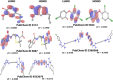
Lassa fever hemorrhagic virus (LFHV) outbreaks in Nigeria continue to increase. With the possibility of a potentially deadly pandemic and the absence of a vaccine, an effective candidate drug that presents with no adverse effects and better clinical outcomes is urgently needed; hence, the aim of the study. Retrieved ligands: 5-(Methylene) evodiamine, N-(2-Fluorobenzoyl) evodiamine, N-[[2-(3-chlorophenyl)-5-(trifluoromethyl) pyrazol-3-yl]methyl]-2-(1-methyl-2,3-dihydroindol-5-yl) propanamide (NCTPMMDP), lauric acid, and sofosbuvir, and the target protein, 3MX5, were prepared and utilised for docking, molecular dynamics simulation, ADMET profiling, and DFT using standard protocols. The docking score for lauric acid was the lowest, with a value of - 6.1, followed by the standard drug and co-crystal control drug with scores of - 9.1 and - 8.9 kcal/mol, respectively. On the other hand, the scores of the other ligands ranged from - 9.5 to - 12.3 kcal/mol. Molecular simulations revealed fluctuations in the RMSD and RMSF values; however, overall, the formed complexes were stable during most of the docking simulation runs. Principal component analysis (PCA) analysis showed that all the ligands, apart from lauric acid, showed better conformational changes within the backbone of the protein. ADMET profiling showed that the ligands were better oral drug candidates than sofosbuvir, as revealed by their better absorption values and absolute compliance with the Lipinski rule of five. The DFT result showed that, apart from lauric acid, all the test ligands possess small energy gaps indicative of high reactivity. Put together, these findings indicate that the test ligands possess anti-LHFV potential that merits further in vivo and in vitro evaluations.
Keywords: DFT; Docking; Lassa fever virus; Nigeria; Nucleoprotein; Simulation.
© 2025. The Author(s).
Conflict of interest statement
Competing interests: The authors declare no competing interests.
Figures


Similar articles
-
Structure-based identification of dual targeting lead inhibitor to Marburg virus glycoprotein and Chandipura virus nucleoprotein: Insights from molecular docking, dynamics and binding free energy analyses.Biochem Biophys Res Commun. 2025 Aug 30;776:152239. doi: 10.1016/j.bbrc.2025.152239. Epub 2025 Jun 20. Biochem Biophys Res Commun. 2025. PMID: 40555000
-
Screening of Propolis compounds reveals potential inhibitors of rhinovirus 3C protease: A computational study.J Mol Graph Model. 2025 Nov;140:109121. doi: 10.1016/j.jmgm.2025.109121. Epub 2025 Jul 3. J Mol Graph Model. 2025. PMID: 40616975
-
Prescription of Controlled Substances: Benefits and Risks.2025 Jul 6. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. 2025 Jul 6. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. PMID: 30726003 Free Books & Documents.
-
Drugs for preventing postoperative nausea and vomiting in adults after general anaesthesia: a network meta-analysis.Cochrane Database Syst Rev. 2020 Oct 19;10(10):CD012859. doi: 10.1002/14651858.CD012859.pub2. Cochrane Database Syst Rev. 2020. PMID: 33075160 Free PMC article.
-
Systemic pharmacological treatments for chronic plaque psoriasis: a network meta-analysis.Cochrane Database Syst Rev. 2021 Apr 19;4(4):CD011535. doi: 10.1002/14651858.CD011535.pub4. Cochrane Database Syst Rev. 2021. Update in: Cochrane Database Syst Rev. 2022 May 23;5:CD011535. doi: 10.1002/14651858.CD011535.pub5. PMID: 33871055 Free PMC article. Updated.
References
-
- Frame, J. D., Baldwin, J. M. Jr, Gocke, D. J. & Troup, J. M. Lassa fever, a new virus disease of man from West Africa. I. Clinical description and pathological findings. Am. J. Trop. Med. Hyg.19(4), 670–676. 10.4269/ajtmh.1970.19.670 (1970). - PubMed
-
- WHO. Lassa fever—Nigeria. (2023, accessed 30 Apr 2024). https://www.who.int/emergencies/disease-outbreak-news/item/2023-DON463.
-
- NCDC. Lassa fever situation report. Epidemiology. Week 7 2024. (2024, accessed 26 Apr 2024). https://ncdc.gov.ng/themes/common/files/sitreps/3702b20687cb7bea4834b4a2....
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

