A single donor cassette enables site-specific knock-in at either the αAmy3 or αAmy8 locus in rice cells via CRISPR/Cas9
- PMID: 40835744
- PMCID: PMC12367948
- DOI: 10.1007/s00253-025-13549-4
A single donor cassette enables site-specific knock-in at either the αAmy3 or αAmy8 locus in rice cells via CRISPR/Cas9
Abstract
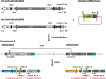
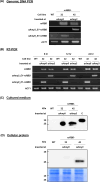
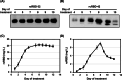
CRISPR/Cas9 gene editing is widely used to manipulate gene expression and integrate transgenes into specific target sites, making it a powerful tool for recombinant protein expression. In this study, we generated a single donor cassette for CRISPR/Cas9-mediated knock-in at either the αAmy3 or αAmy8 locus in rice cells. The transgene was inserted downstream of the promoter and first exon of the rice αAmy3 or αAmy8 genes, which are highly expressed under sugar-starved conditions in rice suspension cultures. We constructed a simple vector with the homologous intron sequences of the αAmy3 and αAmy8, along with rice codon-optimized recombinant receptor binding domain (rcRBD) of the SARS-CoV-2 spike protein, a functional domain responsible for binding to the angiotensin-converting enzyme 2 (ACE2) receptor on host cells. Using this construct, rcRBD was successfully integrated into the intron 1 of either the αAmy3 or αAmy8 genes. As a result, rcRBD expression was driven by endogenous αAmy3 or αAmy8 promoter-signal peptide. Under the control of αAmy3-signal peptide, rcRBD proteins was detected in both the soluble cellular protein fraction and culture medium, whereas expression driven by the αAmy8 promoter-signal peptide was exclusively detected in the culture medium of rice suspension cells. The highest secreted protein yield of rcRBD in the rice culture medium under the control of αAmy8 endogenous promoter reached 20.7 mg/L, demonstrating a production efficiency comparable to that driven by the endogenous αAmy3 promoter.
Keywords: αAmy3 promoter; αAmy8 promoter; CRISPR/Cas9; Intron; Rice suspension cells; Single donor cassette.
© 2025. The Author(s).
Conflict of interest statement
Declarations. Ethical approval: This study did not involve any human participants or animal subjects. All experiments were conducted using rice (O. sativa) cells. Therefore, ethical approval was not required for this research. Competing interests: The authors declare no competing interests.
Figures

Similar articles
-
Applications of CRISPR/Cas9 in a rice protein expression system via an intron-targeted insertion approach.Plant Sci. 2022 Feb;315:111132. doi: 10.1016/j.plantsci.2021.111132. Epub 2021 Nov 26. Plant Sci. 2022. PMID: 35067302
-
High-yield BMP2 expression in rice cells via CRISPR and endogenous αAmy3 promoter.Appl Microbiol Biotechnol. 2024 Feb 14;108(1):206. doi: 10.1007/s00253-024-13054-0. Appl Microbiol Biotechnol. 2024. PMID: 38353738 Free PMC article.
-
Rapid and Robust Generation of Homozygous Fluorescent Reporter Knock-In Cell Pools by CRISPR-Cas9.Cells. 2025 Jul 29;14(15):1165. doi: 10.3390/cells14151165. Cells. 2025. PMID: 40801599 Free PMC article.
-
The effect of sample site and collection procedure on identification of SARS-CoV-2 infection.Cochrane Database Syst Rev. 2024 Dec 16;12(12):CD014780. doi: 10.1002/14651858.CD014780. Cochrane Database Syst Rev. 2024. PMID: 39679851 Free PMC article.
-
Gene editing and CRISPR-dependent homology-mediated end joining.Exp Mol Med. 2025 Jul;57(7):1409-1418. doi: 10.1038/s12276-025-01442-z. Epub 2025 Jul 31. Exp Mol Med. 2025. PMID: 40745005 Free PMC article. Review.
References
-
- Akazawa T, Hara-Nishimura I (1985) Topographic aspects of biosynthesis, extracellular secretion, and intracellular storage of proteins in plant cells. Annu Rev Plant Physiol 36(1):441–472
-
- Arias-Arias JL, Molina-Castro SE, Monturiol-Gross L, Lomonte B, Corrales-Aguilar E (2023) Stable production of recombinant SARS-CoV-2 receptor-binding domain in mammalian cells with co-expression of a fluorescent reporter and its validation as antigenic target for COVID-19 serology testing. Biotechnology Reports 37:e00780 - PMC - PubMed
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

