Highly accurate prophage island detection with PIDE
- PMID: 40836306
- PMCID: PMC12366036
- DOI: 10.1186/s13059-025-03733-0
Highly accurate prophage island detection with PIDE
Abstract
As important mobile elements in prokaryotes, prophages shape the genomic context of their hosts and regulate the structure of bacterial populations. However, it is challenging to precisely identify prophages through computational methods. Here, we introduce PIDE for identifying prophages from bacterial genomes or metagenome-assembled genomes. PIDE integrates a pre-trained protein language model and gene density clustering algorithm to distinguish prophages. Benchmarking with induced prophage sequencing datasets demonstrates that PIDE pinpoints prophages with precise boundaries. Applying PIDE to 4744 human gut representative genomes reveals 24,467 prophages with widespread functional capacity. PIDE is available at https://github.com/chyghy/PIDE , with model training code at https://zenodo.org/records/16457629 .
Keywords: Gene cluster; Human gut metagenome; Prophage identification; Protein language model.
© 2025. The Author(s).
Conflict of interest statement
Declarations. Ethics approval and consent to participate: Not applicable. Consent for publication: Not applicable. Competing interests: The authors declare no competing interests.
Figures


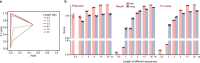

References
-
- Liang G, Zhao C, Zhang H, Mattei L, Sherrill-Mix S, Bittinger K, et al. The stepwise assembly of the neonatal virome is modulated by breastfeeding. Nat. 2020;581(7809):470–4. http://www.nature.com/articles/s41586-020-2192-1 - PMC - PubMed
MeSH terms
LinkOut - more resources
Full Text Sources
Research Materials

