Individual-based modeling unravels spatial and social interactions in bacterial communities
- PMID: 40838740
- PMCID: PMC12411854
- DOI: 10.1093/ismejo/wraf116
Individual-based modeling unravels spatial and social interactions in bacterial communities
Abstract
Bacterial interactions are fundamental in shaping community structure and function, driving processes that range from plastic degradation in marine ecosystems to dynamics within the human gut microbiome. Yet, studying these interactions is challenging due to difficulties in resolving spatiotemporal scales, quantifying interaction strengths, and integrating intrinsic cellular behaviors with extrinsic environmental conditions. Individual-based modeling addresses these challenges through single-cell-level simulations that explicitly model growth, division, motility, and environmental responses. By capturing both the spatial organization and social interactions, individual-based modeling reveals how microbial interactions and environmental gradients collectively shape community architecture, species coexistence, and adaptive responses. In particular, individual-based modeling provides mechanistic insights into how social behaviors-such as competition, metabolic cooperation, and quorum sensing-are regulated by spatial structure, uncovering the interplay between localized interactions and emergent community properties. In this review, we synthesize recent applications of individual-based modeling in studying bacterial spatial and social interactions, highlighting how their interplay governs community stability, diversity, and resilience. By linking individual-scale interactions with the ecosystem-level organization, individual-based modeling offers a predictive framework for understanding microbial ecology and informing strategies for controlling and engineering bacterial consortia in both natural and applied settings.
Keywords: adaptive strategies; individual variability; individual-based modeling (IbM); metabolic cooperation; niche differentiation; resource gradient; social interactions; spatial heterogeneity; spatial interactions.
© The Author(s) 2025. Published by Oxford University Press on behalf of the International Society for Microbial Ecology.
Conflict of interest statement
None declared.
Figures
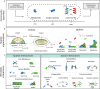



References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

