Leveraging Retrieval-Augmented Large Language Models for Dietary Recommendations With Traditional Chinese Medicine's Medicine Food Homology: Algorithm Development and Validation
- PMID: 40840437
- PMCID: PMC12370266
- DOI: 10.2196/75279
Leveraging Retrieval-Augmented Large Language Models for Dietary Recommendations With Traditional Chinese Medicine's Medicine Food Homology: Algorithm Development and Validation
Abstract
Background: Traditional Chinese Medicine (TCM) emphasizes the concept of medicine food homology (MFH), which integrates dietary therapy into health care. However, the practical application of MFH principles relies heavily on expert knowledge and manual interpretation, posing challenges for automating MFH-based dietary recommendations. Although large language models (LLMs) have shown potential in health care decision support, their performance in specialized domains such as TCM is often hindered by hallucinations and a lack of domain knowledge. The integration of uncertain knowledge graphs (UKGs) with LLMs via retrieval-augmented generation (RAG) offers a promising solution to overcome these limitations by enabling a structured and faithful representation of MFH principles while enhancing LLMs' ability to understand the inherent uncertainty and heterogeneity of TCM knowledge. Consequently, it holds potential to improve the reliability and accuracy of MFH-based dietary recommendations generated by LLMs.
Objective: This study aimed to introduce Yaoshi-RAG, a framework that leverages UKGs to enhance LLMs' capabilities in generating accurate and personalized MFH-based dietary recommendations.
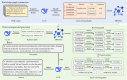
Methods: The proposed framework began by constructing a comprehensive MFH knowledge graph (KG) through LLM-driven open information extraction, which extracted structured knowledge from multiple sources. To address the incompleteness and uncertainty within the MFH KG, UKG reasoning was used to measure the confidence of existing triples and to complete missing triples. When processing user queries, query entities were identified and linked to the MFH KG, enabling retrieval of relevant reasoning paths. These reasoning paths were then ranked based on triple confidence scores and entity importance. Finally, the most informative reasoning paths were encoded into prompts using prompt engineering, enabling the LLM to generate personalized dietary recommendations that aligned with both individual health needs and MFH principles. The effectiveness of Yaoshi-RAG was evaluated through both automated metrics and human evaluation.
Results: The constructed MFH KG comprised 24,984 entities, 22 relations, and 29,292 triples. Extensive experiments demonstrate the superiority of Yaoshi-RAG in different evaluation metrics. Integrating the MFH KG significantly improved the performance of LLMs, yielding an average increase of 14.5% in Hits@1 and 8.7% in F1-score, respectively. Among the evaluated LLMs, DeepSeek-R1 achieved the best performance, with 84.2% in Hits@1 and 71.5% in F1-score, respectively. Human evaluation further validated these results, confirming that Yaoshi-RAG consistently outperformed baseline models across all assessed quality dimensions.
Conclusions: This study shows Yaoshi-RAG, a new framework that enhances LLMs' capabilities in generating MFH-based dietary recommendations through the knowledge retrieved from a UKG. By constructing a comprehensive TCM knowledge representation, our framework effectively extracts and uses MFH principles. Experimental results demonstrate the effectiveness of our framework in synthesizing traditional wisdom with advanced language models, facilitating personalized dietary recommendations that address individual health conditions while providing evidence-based explanations.
Keywords: Traditional Chinese Medicine; dietary recommendation; large language model; medicine food homology; retrieval-augmented generation; uncertain knowledge graph.
© Hangyu Sha, Fan Gong, Bo Liu, Runfeng Liu, Haofen Wang, Tianxing Wu. Originally published in JMIR Medical Informatics (https://medinform.jmir.org).
Conflict of interest statement
Figures

Similar articles
-
Enhancing Pulmonary Disease Prediction Using Large Language Models With Feature Summarization and Hybrid Retrieval-Augmented Generation: Multicenter Methodological Study Based on Radiology Report.J Med Internet Res. 2025 Jun 11;27:e72638. doi: 10.2196/72638. J Med Internet Res. 2025. PMID: 40499132 Free PMC article.
-
Evaluating and Improving Syndrome Differentiation Thinking Ability in Large Language Models: Method Development Study.JMIR Med Inform. 2025 Jun 20;13:e75103. doi: 10.2196/75103. JMIR Med Inform. 2025. PMID: 40540614 Free PMC article.
-
Prescription of Controlled Substances: Benefits and Risks.2025 Jul 6. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. 2025 Jul 6. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. PMID: 30726003 Free Books & Documents.
-
Stench of Errors or the Shine of Potential: The Challenge of (Ir)Responsible Use of ChatGPT in Speech-Language Pathology.Int J Lang Commun Disord. 2025 Jul-Aug;60(4):e70088. doi: 10.1111/1460-6984.70088. Int J Lang Commun Disord. 2025. PMID: 40627744 Review.
-
Cost-effectiveness of using prognostic information to select women with breast cancer for adjuvant systemic therapy.Health Technol Assess. 2006 Sep;10(34):iii-iv, ix-xi, 1-204. doi: 10.3310/hta10340. Health Technol Assess. 2006. PMID: 16959170
References
-
- Zhong H, Tang ZQ, Li YF, Wang M, Sun WY, He RR. The evolution and significance of medicine and food homology. Acupunct Herbal Med. 2024;4(1):19–35. doi: 10.1097/HM9.0000000000000104. doi. - DOI
-
- Zhao W, Zhou K, Li J, et al. A survey of large language models. arXiv. 2023 doi: 10.48550/arXiv.2303.18223. Preprint posted online on. doi. - DOI
MeSH terms
LinkOut - more resources
Full Text Sources

