An alternative cytoplasmic SFPQ isoform with reduced phase separation potential is up-regulated in ALS
- PMID: 40845103
- PMCID: PMC12372870
- DOI: 10.1126/sciadv.adt4814
An alternative cytoplasmic SFPQ isoform with reduced phase separation potential is up-regulated in ALS
Abstract
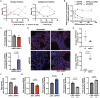
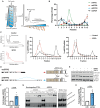
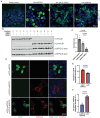
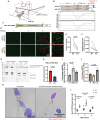
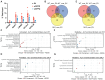
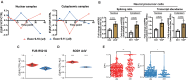
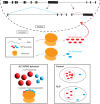
Splicing factor proline- and glutamine-rich (SFPQ) is an RNA binding protein that broadly regulates RNA metabolism. Although its nuclear roles are well studied, evidence of SFPQ's cytoplasmic functionality is emerging. Altered expression and nuclear-to-cytoplasmic redistribution of SFPQ have been recognized in amyotrophic lateral sclerosis (ALS) pathology, yet the mechanistic bases for these phenomena remain undetermined. We identified altered SFPQ splicing in ALS, increasing the expression of an alternative mRNA isoform lacking a nuclear localization sequence, which we termed "altSFPQ." We find that altSFPQ mRNA contributes to SFPQ autoregulation and is highly unstable yet exhibits context-specific translation with cytoplasm-predominant localization. Notably, reduced canonical SFPQ coincides with increased altSFPQ transcript expression in familial and sporadic ALS models, providing a mechanistic basis for SFPQ nuclear-to-cytoplasmic redistribution in patients with ALS. Last, we observe that the altSFPQ protein has reduced phase separation potential and differential protein binding compared to its canonical counterpart, providing insight into its mechanistic relevance to physiology and ALS pathogenesis.
Figures
Similar articles
-
Structural dynamics of IDR interactions in human SFPQ and implications for liquid-liquid phase separation.Acta Crystallogr D Struct Biol. 2025 Jul 1;81(Pt 7):357-379. doi: 10.1107/S2059798325005303. Epub 2025 Jun 27. Acta Crystallogr D Struct Biol. 2025. PMID: 40574713 Free PMC article.
-
A novel nuclear RNA HSD52 scaffolding NONO/SFPQ complex modulates DNA damage repair to facilitate temozolomide resistance.Neuro Oncol. 2025 May 15;27(4):963-978. doi: 10.1093/neuonc/noae272. Neuro Oncol. 2025. PMID: 39673809
-
Deep learning analyses of splicing variants identify the link of PCP4 with amyotrophic lateral sclerosis.Brain. 2025 Jul 7;148(7):2331-2347. doi: 10.1093/brain/awaf025. Brain. 2025. PMID: 39852553
-
The Emerging Role of the RNA-Binding Protein SFPQ in Neuronal Function and Neurodegeneration.Int J Mol Sci. 2020 Sep 28;21(19):7151. doi: 10.3390/ijms21197151. Int J Mol Sci. 2020. PMID: 32998269 Free PMC article. Review.
-
Alternative Splicing of Exon 23a in Neurofibromatosis Type 1 Pre-mRNA: Its Contribution to the Protein Structure and Function of Neurofibromin.Wiley Interdiscip Rev RNA. 2025 Jul-Aug;16(4):e70021. doi: 10.1002/wrna.70021. Wiley Interdiscip Rev RNA. 2025. PMID: 40812791 Free PMC article. Review.
References
-
- Neumann M., Sampathu D. M., Kwong L. K., Truax A. C., Micsenyi M. C., Chou T. T., Bruce J., Schuck T., Grossman M., Clark C. M., McCluskey L. F., Miller B. L., Masliah E., Mackenzie I. R., Feldman H., Feiden W., Kretzschmar H. A., Trojanowski J. Q., Lee V. M., Ubiquitinated TDP-43 in frontotemporal lobar degeneration and amyotrophic lateral sclerosis. Science 314, 130–133 (2006). - PubMed
-
- Kwiatkowski T. J. Jr., Bosco D. A., Leclerc A. L., Tamrazian E., Vanderburg C. R., Russ C., Davis A., Gilchrist J., Kasarskis E. J., Munsat T., Valdmanis P., Rouleau G. A., Hosler B. A., Cortelli P., de Jong P. J., Yoshinaga Y., Haines J. L., Pericak-Vance M. A., Yan J., Ticozzi N., Siddique T., McKenna-Yasek D., Sapp P. C., Horvitz H. R., Landers J. E., Brown R. H. Jr., Mutations in the FUS/TLS gene on chromosome 16 cause familial amyotrophic lateral sclerosis. Science 323, 1205–1208 (2009). - PubMed
-
- Vance C., Rogelj B., Hortobágyi T., De Vos K. J., Nishimura A. L., Sreedharan J., Hu X., Smith B., Ruddy D., Wright P., Ganesalingam J., Williams K. L., Tripathi V., Al-Saraj S., Al-Chalabi A., Leigh P. N., Blair I. P., Nicholson G., de Belleroche J., Gallo J.-M., Miller C. C., Shaw C. E., Mutations in FUS, an RNA processing protein, cause familial amyotrophic lateral sclerosis type 6. Science 323, 1208–1211 (2009). - PMC - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

