Plasma exosomes from individuals with type 2 diabetes drive breast cancer aggression in patient-derived organoids
- PMID: 40858992
- PMCID: PMC12381303
- DOI: 10.1038/s42003-025-08663-y
Plasma exosomes from individuals with type 2 diabetes drive breast cancer aggression in patient-derived organoids
Abstract
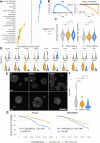
Women with obesity-driven type 2 diabetes (T2D) face worse breast cancer outcomes, yet metabolic status does not fully inform current standards of care. We previously identified plasma exosomes as key drivers of tumor progression; however, their effect on immune cells within the tumor microenvironment (TME) remains unclear. Using a novel patient-derived organoid (PDO) system that preserves native tumor-infiltrating lymphocytes (TILs), we show that T2D plasma exosomes induce a 13.6-fold expansion of immunosuppressive TILs relative to nondiabetic controls. This immune dysfunction may promote micrometastatic survival and resistance to checkpoint blockade, a known issue in T2D cancer patients. Tumor-intrinsic analysis revealed a 1.5-fold increase in intratumoral heterogeneity and 2.3-fold upregulation of aggressive signaling networks. These findings reveal how T2D-associated metabolic dysregulation alters tumor-immune crosstalk through previously underappreciated exosomal signaling, impairing antitumor immunity and accelerating progression. Understanding these dynamics could inform tailored therapies for this high-risk, underserved patient population.
© 2025. The Author(s).
Conflict of interest statement
Competing interests: The authors declare no competing interests.
Figures






Update of
-
Plasma exosomes from individuals with type 2 diabetes drive breast cancer aggression in patient-derived organoids.bioRxiv [Preprint]. 2025 Apr 30:2024.09.13.612950. doi: 10.1101/2024.09.13.612950. bioRxiv. 2025. Update in: Commun Biol. 2025 Aug 26;8(1):1276. doi: 10.1038/s42003-025-08663-y. PMID: 39345362 Free PMC article. Updated. Preprint.
References
MeSH terms
Grants and funding
- R01CA222170/U.S. Department of Health & Human Services | NIH | National Cancer Institute (NCI)
- U01 CA243004/CA/NCI NIH HHS/United States
- U01 CA182898/CA/NCI NIH HHS/United States
- 5T32AI007309/U.S. Department of Health & Human Services | NIH | National Institute of Allergy and Infectious Diseases (NIAID)
- T32 AI007309/AI/NIAID NIH HHS/United States
LinkOut - more resources
Full Text Sources
Medical

