KLK1 as an Epithelial-Specific Brake Inhibits Colorectal Tumorigenesis by Suppressing B1R-Mediated Fibroblast Phenotypic Transition
- PMID: 40859429
- PMCID: PMC12622486
- DOI: 10.1002/advs.202507063
KLK1 as an Epithelial-Specific Brake Inhibits Colorectal Tumorigenesis by Suppressing B1R-Mediated Fibroblast Phenotypic Transition
Abstract
Inflammatory bowel disease (IBD) is increasing worldwide, and the persistence of chronic inflammation may lead to colitis-associated colorectal cancer (CAC). KLK1 expression is reduced in colitis, and its potential role in the intestinal mucosal barrier is still unclear. Here, KLK1 is investigated whether a supplement can reduce colitis and colorectal carcinogenesis. This study investigated KLK1's protective function in intestinal barrier integrity using Dextran Sulfate Sodium Salt (DSS) / Azoxymethane (AOM)-DSS-induced colitis/CAC models, Apc-deficient mice, and human clinical samples. KLK1-AAV2 knockdown mice exhibited exacerbated colitis symptoms, including severe diarrhea and impaired mucosal barrier markers, while KLK1 levels are notably reduced in ulcerative colitis patients and colorectal cancer specimens. Mechanistically, bradykinin receptor B1 (B1R) upregulation in CAC models activated extracellular matrix pathways, driving fibroblast phenotypic shifts that disrupt stromal homeostasis. Crucially, KLK1 supplementation reversed these pathological changes, demonstrating its dual role in maintaining epithelial barrier function and regulating fibroblast-ECM interactions. These findings position KLK1 as a potential therapeutic target for colitis and CRC chemoprevention, offering novel insights into IBD pathogenesis through its modulation of mucosal protection and stromal remodeling processes.
Keywords: bradykinin B1 receptors; cancer‐associated fibroblasts; extracellular matrix; intestinal barrier; tissue kallikrein.
© 2025 The Author(s). Advanced Science published by Wiley‐VCH GmbH.
Conflict of interest statement
The authors declare no conflict of interest.
Figures





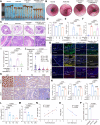




References
-
- Ordás I., Eckmann L., Talamini M., Baumgart D. C., Sandborn W. J., Lancet 2012, 380, 1606. - PubMed
-
- Lundwall Å., Band V., Blaber M., Clements J. A., Courty Y., Diamandis E. P., Fritz H., Lilja H., Malm J., Maltais L. J., Yvonne Olsson A., Petraki C., Scorilas A., Sotiropoulou G., Stenman U.‐H., Stephan C., Talieri M., Yousef G. M., Biol. Chem. 2006, 387, 637. - PubMed
-
- Prassas I., Eissa A., Poda G., Diamandis E. P., Nat. Rev. Drug Discovery 2015, 14, 183. - PubMed
-
- Marceau F., Regoli D., Nat. Rev. Drug Discovery 2004, 3, 845. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
