The Chelating Abilities of Tertiary Amines with N-O-Donors Towards Cu(II) Ions and the Catalytic Properties of the Resulting Complexes
- PMID: 40871572
- PMCID: PMC12388300
- DOI: 10.3390/molecules30163419
The Chelating Abilities of Tertiary Amines with N-O-Donors Towards Cu(II) Ions and the Catalytic Properties of the Resulting Complexes
Abstract
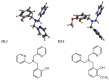
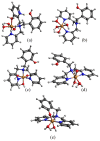
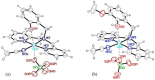
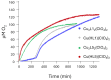
Oxidative stress, driven by excess reactive oxygen species (ROS), is a key factor in the progression of neurodegenerative diseases like Alzheimer's disease (AD). In this context, copper dysregulation can also contribute to this imbalance, being responsible for enhanced ROS production, so that copper scavenging has been investigated as a possible therapeutic strategy. This study investigates the behavior of two isostructural ligands, featuring an N3O donor set, that effectively chelate Cu(II) in aqueous solution. Interestingly, their resulting mono- or dinuclear copper complexes feature a coordination environment suitable to foster antioxidant activity. By transforming copper's oxidant potential into antioxidant action, these systems may reduce copper-induced oxidative damage. The work examines the pH-dependent metal-binding behavior of the ligands, the catalytic properties of the resulting complexes under physiological conditions, and their ability to inhibit β-amyloid peptide aggregation.
Keywords: N3O donors; amyloid aggregation; antioxidants; chelators; copper complexes.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures




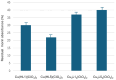
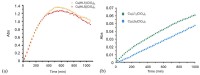

References
-
- Jomova K., Alomar S.Y., Alwasel S.H., Nepovimova E., Kuca K., Valko M. Several lines of antioxidant defense against oxidative stress: Antioxidant enzymes, nanomaterials with multiple enzyme-mimicking activities, and low-molecular-weight antioxidants. Arch Toxicol. 2024;98:1323–1367. doi: 10.1007/s00204-024-03696-4. - DOI - PMC - PubMed
-
- López M.B., Oterino M.B., González J.M. The Structural Biology of Catalase Evolution. Subcell Biochem. 2024;104:33–47. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

