N-[(Thiophen-3-yl)methyl]benzamides as Fusion Inhibitors of Influenza Virus Targeting H1 and H5 Hemagglutinins
- PMID: 40880513
- PMCID: PMC12434670
- DOI: 10.1021/acs.jmedchem.5c01357
N-[(Thiophen-3-yl)methyl]benzamides as Fusion Inhibitors of Influenza Virus Targeting H1 and H5 Hemagglutinins
Abstract
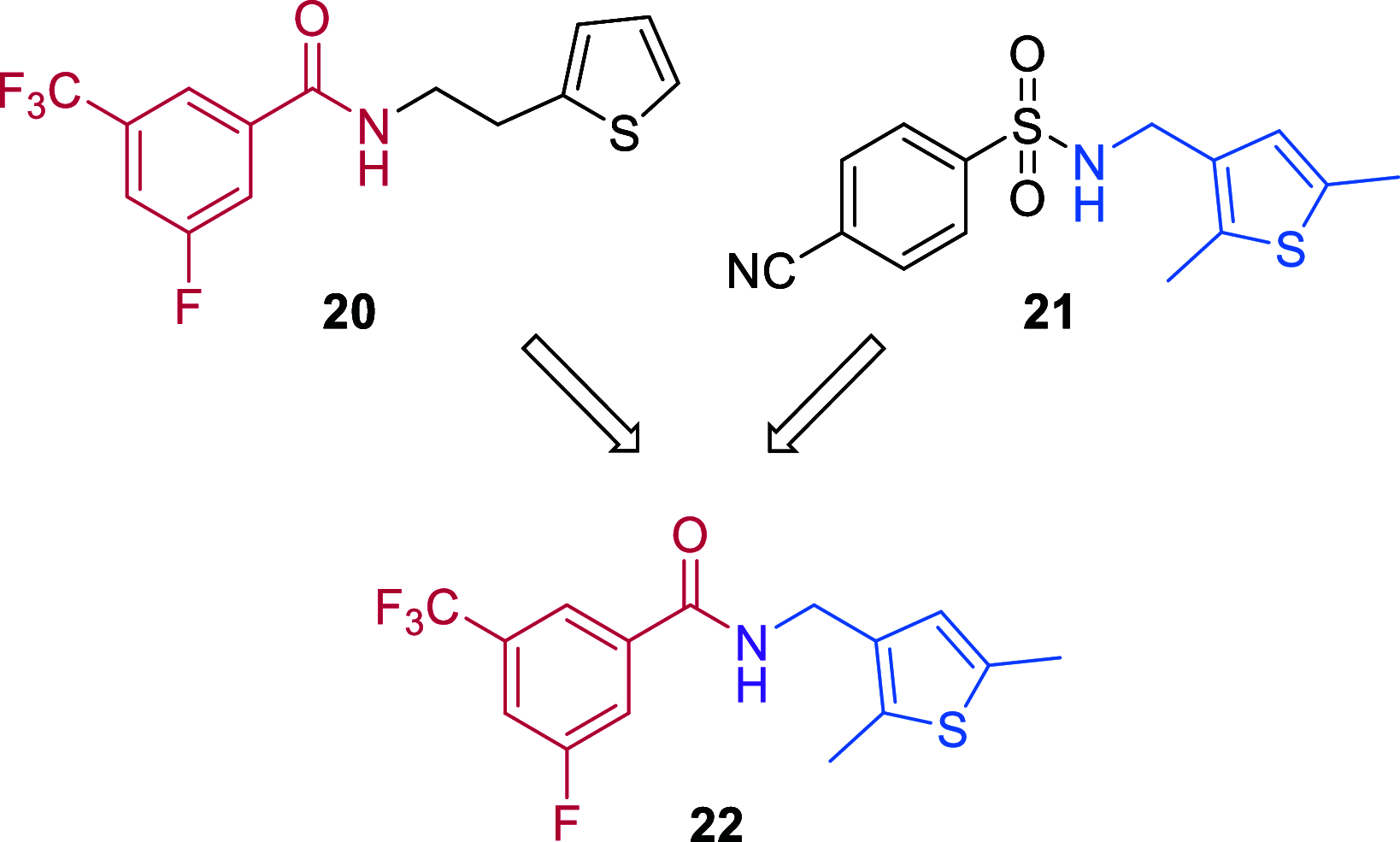
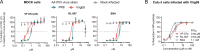
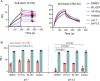
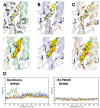
Novel antiviral drugs are needed to prepare for infections from influenza A virus (IAV). Here, a series of N-[(thiophen-3-yl)methyl]benzamides, which target the hemagglutinin (HA)-mediated fusion process, is reported. The most active compound, VF-57a, displays a 50% effective concentration (EC50) of ∼0.8 μM and an antiviral selectivity index >130 in Madin-Darby canine kidney (MDCK) cells infected with A/H1N1 virus. VF-57a proved to be a strong inhibitor of A/H1N1 and A/H5N1 pseudovirus entry (EC50 values of 0.3 and 0.8 μM, respectively). Cell-cell fusion assays in HA-expressing cells, surface plasmon resonance-based assessment of HA protein refolding, and resistance studies suggested that VF-57a prevents the conformational change of HA at acidic pH. Molecular modeling highlighted the role of the dimethylthiophene moiety and the amide-based tether in anchoring to the binding cavity of HA. Our findings support the further development of this class of IAV fusion inhibitors against A/H1N1 and A/H5N1 viruses.
Figures






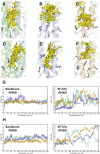



References
-
- Iuliano A. D., Roguski K. M., Chang H. H., Muscatello D. J., Palekar R., Tempia S., Cohen C., Gran J. M., Schanzer D., Cowling B. J.. et al. Estimates of global seasonal influenza-associated respiratory mortality: A modelling study. Lancet. 2018;391(10127):1285–1300. doi: 10.1016/s0140-6736(17)33293-2. - DOI - PMC - PubMed
-
- Dawood F. S., Iuliano A. D., Reed C., Meltzer M. I., Shay D. K., Cheng P.-Y., Bandaranayake D., Breiman R. F., Brooks W. A., Buchy P., Feikin D. R., Fowler K. B., Gordon A., Hien N. T., Horby P., Huang Q. S., Katz M. A., Krishnan A., Lal R., Montgomery J. M., Mølbak K., Pebody R., Presanis A. M., Razuri H., Steens A., Tinoco Y. O., Wallinga J., Yu H., Vong S., Bresee J., Widdowson M.-A.. Estimated global mortality associated with the first 12 months of 2009 pandemic influenza A H1N1 virus circulation: a modelling study. Lancet Infect. Dis. 2012;12(9):687–695. doi: 10.1016/S1473-3099(12)70121-4. - DOI - PubMed
-
- Nguyen T. Q., Hutter C. R., Markin A., Thomas M., Lantz K., Killian M. L., Janzen G. M., Vijendran S., Wagle S., Inderski B., Magstadt D. R., Li G., Diel D. G., Frye E. A., Dimitrov K. M., Swinford A. K., Thompson A. C., Snekvik K. R., Suarez D. L., Lakin S. M., Schwabenlander S., Ahola S. C., Johnson K. R., Baker A. L., Robbe-Austerman S., Torchetti M. K., Anderson T. K.. Emergence and interstate spread of highly pathogenic avian influenza A(H5N1) in dairy cattle in the United States. Science. 2025;388(6745):eadq0900. doi: 10.1126/science.adq0900. - DOI - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical

