A tutorial for mesoscale computer simulations of lipid membranes: tether pulling, tubulation and fluctuations
- PMID: 40905106
- PMCID: PMC12409609
- DOI: 10.1039/d5sm00148j
A tutorial for mesoscale computer simulations of lipid membranes: tether pulling, tubulation and fluctuations
Abstract
Lipid membranes and membrane deformations are a long-standing area of research in soft matter and biophysics. Computer simulations have complemented analytical and experimental approaches as one of the pillars in the field. However, setting up and using membrane simulations can come with barriers due to the multidisciplinary effort involved and the vast choice of existing simulations models. In this review, we introduce the non-expert reader to coarse-grained membrane simulations at the mesoscale. Firstly, we give a concise overview of the modelling approaches to study fluid membranes, together with guidance to more specialized references. Secondly, we provide a conceptual guide on how to develop mesoscale membrane simulations. Lastly, we construct a hands-on tutorial on how to apply mesoscale membrane simulations, by providing a pedagogical examination of membrane tether pulling, shape and mechanics of membrane tubes, and membrane fluctuations with three different membrane models, and discussing them in terms of their scope and how resource-intensive they are. To ease the reader's venture into the field, we provide a repository with ready-to-run tutorials.
Conflict of interest statement
There are no conflicts to declare.
Figures

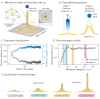



References
-
- Alberts B., Johnson A., Lewis J., Morgan D., Raff M., Roberts K. and Walter P., Molecular biology of the cell, Garland Science, New York, 6th edn, 2015
-
- Physics of Biological Membranes, ed. P. Bassereau and P. Sens, Springer International Publishing, Cham, 2018
-
- Phillips R., Kondev J., Theriot J. and Garcia H., Physical biology of the cell, Garland Science, New York, 2012
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

