Circulating extracellular vesicle isomiR signatures predict therapy response in patients with multiple myeloma
- PMID: 40961943
- PMCID: PMC12629820
- DOI: 10.1016/j.xcrm.2025.102358
Circulating extracellular vesicle isomiR signatures predict therapy response in patients with multiple myeloma
Abstract
Multiple myeloma (MM) is a plasma cell neoplasm characterized by high inter- and intra-patient clonal heterogeneity, leading to high variability in therapeutic responses. Minimally invasive biomarkers that predict response may help personalize treatment decisions. IsoSeek, a single-nucleotide resolution small RNA sequencing method can profile thousands of microRNAs (miRNAs) and their variants (isomiRs) from patient plasma-purified extracellular vesicles (EVs). Machine learning-generated miRNA/isomiR classifiers accurately predict therapeutic response in relapsed/refractory MM (RRMM) patients receiving daratumumab-containing regimens, achieving an area-under-the-curve of 0.98 (95% confidence interval [CI]:0.94-1.00). A classifier signature with the plasma cell-selective miR-148-3p, predicts durable response (≥6 months), progression-free (hazard ratio [HR]: 33.09, 95% CI: 4.2-262, p < 0.001), and overall survival (HR: 3.81, 95% CI: 1.05-13.99, p < 0.05). Targetome analysis connects the prognostic classifier to established MM drug targets BCL2 and MYC suggesting biological relevance. Thus, EV-isomiR sequencing in MM patients offers a tumor-naïve alternative to an invasive bone-marrow biopsy for predicting treatment outcome.
Keywords: extracellular vesicles; isomiR modelling; liquid biopsy; miRNAs; multiple myeloma; personalized therapy; response prediction.
Copyright © 2025 The Author(s). Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests D.M.P. and M.H. were co-founders of Exbiome BV. D.M.P. was CSO of ExBiome BV and served as an advisor for Takeda for which he received travel compensation. D.M.P. received research funding from Gilead, AbbVie (not related to this project), and Amgen (related to this project). C.G.-M. and M.A.J.v.E. received travel compensation from QIAGEN. ExBiome received funding from Amgen for sequencing the samples. Amgen had no role in design of the study and was not involved in the writing of this manuscript. NWCJvdD has received research support from Janssen Pharmaceuticals, Amgen, Celgene, Novartis, Cellectis, and BMS and serves in advisory boards for Janssen Pharmaceuticals, Amgen, Celgene, BMS, Sanofi, Takeda, Roche, Novartis, Bayer, Adaptive, Merck, Kite Pharma, Pfizer, AbbVie, and Servier, all paid to institution.
Figures




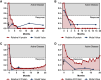

References
-
- San-Miguel J., Avet-Loiseau H., Paiva B., Kumar S., Dimopoulos M.A., Facon T., Mateos M.-V., Touzeau C., Jakubowiak A., Usmani S.Z., et al. Sustained minimal residual disease negativity in newly diagnosed multiple myeloma and the impact of daratumumab in MAIA and ALCYONE. Blood. 2022;139:492–501. doi: 10.1182/blood.2020010439. - DOI - PMC - PubMed
-
- Larrayoz M., Garcia-Barchino M.J., Celay J., Etxebeste A., Jimenez M., Perez C., Ordoñez R., Cobaleda C., Botta C., Fresquet V., et al. Preclinical models for prediction of immunotherapy outcomes and immune evasion mechanisms in genetically heterogeneous multiple myeloma. Nat. Med. 2023;29:632–645. doi: 10.1038/s41591-022-02178-3. - DOI - PMC - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical

