Proteomic analysis uncovers biological diversity in molecularly defined endometrial carcinomas
- PMID: 40966993
- PMCID: PMC12478120
- DOI: 10.1016/j.neo.2025.101229
Proteomic analysis uncovers biological diversity in molecularly defined endometrial carcinomas
Abstract
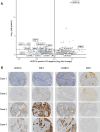
While endometrial cancer has an overall favorable prognosis, some patients have poor outcomes and may benefit from further refinements of the current classification systems. Molecular classification stratifies endometrial cancer patients into four prognostic subtypes: POLEmut, MMRd (mismatch repair deficient), p53abn, and NSMP (no specific molecular profile), where patients with POLEmut have the best prognosis and p53abn has the worst prognosis. We used proteomic profiling to assess if additional prognostic or predictive information could be identified across or within molecular subtypes. Global proteome profiling of formalin fixed, paraffin embedded samples, that had clinicopathologic and outcome data, was performed on 184 endometrial cancers encompassing all four molecular subtypes, including replicate samples of the same tumor, and both biopsy and final hysterectomy specimens. To ensure representation of each subtype, we profiled an approximately equal distribution in the 148 unique tumors; 34 (23%) POLEmut, 40 (27%) MMRd, 35 (24%) p53abn and 39 (26%) NSMP, rather than the population-based distributions. There was high reproducibility in the proteomic profiles of intra-tumor replicate samples, and between matched biopsy and hysterectomy tumor samples. Consensus clustering identified four clusters with different prognosis, named 'Adhesion', 'Immune', 'Proliferation', and 'Metabolic' based on the functional characteristics of the enriched proteins. We associated protein expression features with common mutations, molecular subtype, and outcomes. These results demonstrate the biologic diversity within endometrial cancers, both between and within molecular subtypes, and provide candidate features for functional and clinical investigation.
Keywords: Endometrial carcinoma; Molecular subtype; Proteomics.
Copyright © 2025. Published by Elsevier Inc.
Conflict of interest statement
Declaration of competing interest The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Figures




References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

