Swine influenza-modified pulmonary microbiota
- PMID: 40989185
- PMCID: PMC12450909
- DOI: 10.3389/fcimb.2025.1634469
Swine influenza-modified pulmonary microbiota
Abstract
Influenza A virus (IAV) remains a major health concern in both humans and animals, with pigs serving as key reservoirs for generating novel reassortant viruses with pandemic potential. Respiratory microbiome alterations during infection may facilitate secondary bacterial complications. This study investigates the lung microbiota of pigs naturally infected with IAV across different regions in Spain, using Oxford Nanopore Technologies (ONT) long-read 16S rRNA sequencing to characterize associated bacterial communities. Our results show a higher bacterial genus diversity in IAV-infected animals compared to healthy controls, with significant differences in both presence and relative abundance of bacterial taxa. Infected lungs exhibited increased proportions of potential pathogens, particularly Glaesserella spp., detected in approximately 60% of infected samples, often as the dominant genus. Other pathogenic genera, including Pasteurella, Staphylococcus, Mycoplasma, and Fusobacterium, were also strongly associated with infection. Clustering analyses revealed distinct microbial profiles that clearly separated infected from non-infected animals, identifying specific bacterial signatures predictive of infection status. These findings suggest that IAV infection significantly alters the pulmonary microbiota, potentially creating a permissive environment for secondary bacterial infections. This study underscores the relevance of microbiota shifts during IAV infection in swine and highlights the importance of understanding microbial dynamics in respiratory disease progression. Additionally, we present a novel, rapid, and practical experimental pipeline based on ONT long-read sequencing to investigate the respiratory microbiota in swine infection models. This approach offers a valuable tool for future research and potential diagnostic applications in both veterinary and human medicine.
Keywords: Oxford Nanopore; coinfection; influenza virus; lung; pigs; respiratory microbiome; sequencing; swine.
Copyright © 2025 Arranz-Herrero, Izpura-Luis, Presa, Reche, Encinas, Kwon, Rius-Rocabert, Tur-Planells, Tejerina, Ochando, Gutiérrez-Martín, Bortz, Garcia-Sastre, Richt, Montoya, del Real and Nistal-Villan.
Conflict of interest statement
The A.G.-S. laboratory has received research support from Avimex, Dynavax, Pharmamar, 7Hills Pharma, ImmunityBio and Accurius, outside of the reported work. A.G.-S. has consulting agreements for the following companies involving cash and/or stock: Castlevax, Amovir, Vivaldi Biosciences, Contrafect, 7Hills Pharma, Avimex, Pagoda, Accurius, Esperovax, Applied Biological Laboratories, Pharmamar, CureLab Oncology, CureLab Veterinary, Synairgen, Paratus, Pfizer, Virofend and Prosetta, outside of the reported work. A.G.-S. has been an invited speaker in meeting events organized by Seqirus, Janssen, Abbott, Astrazeneca and Novavax. A.G.-S. is inventor on patents and patent applications on the use of antivirals and vaccines for the treatment and prevention of virus infections and cancer, owned by the Icahn School of Medicine at Mount Sinai, New York, outside of the reported work. The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures


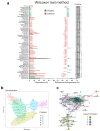
References
-
- Cáceres M. D., Jansen F., Endicott S., Dell N. (2025).
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

