SMAD4 induces opposite effects on metastatic growth from pancreatic tumors depending on the organ of residence
- PMID: 40999053
- PMCID: PMC12643927
- DOI: 10.1038/s43018-025-01047-5
SMAD4 induces opposite effects on metastatic growth from pancreatic tumors depending on the organ of residence
Abstract
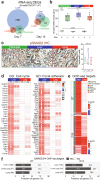
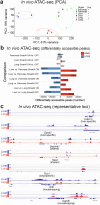
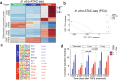
The role of driver gene mutations in sustaining tumor growth at metastatic sites is poorly understood. SMAD4 inactivation is a paradigm of such mutations and a hallmark of pancreatic ductal adenocarcinoma (PDAC). To determine whether metastatic tumors are dependent on SMAD4 inactivation, we developed a mouse model of PDAC that enables spatiotemporal control of Smad4 expression. While Smad4 inactivation in the premalignant pancreas facilitated the formation of primary tumors, Smad4 reactivation in metastatic disease suppressed liver metastases but promoted lung metastases. These divergent effects were underpinned by organ-biased differences in the tumor cells' chromatin state that emerged in the premalignant pancreas and were distinguished by the dominance of KLF4 versus RUNX1 transcription factors. Our results show how epigenetic states favored by the organ of residence can influence the output of driver mutations in metastatic tumors, which has implications for interpreting tumor genetics and therapeutically targeting metastatic disease.
© 2025. The Author(s).
Conflict of interest statement
Competing interests: S.W.L. is a consultant for Fate Therapeutics and is a consultant and holds equity in Blueprint Medicines, ORIC Pharmaceuticals, Mirimus, PMV Pharmaceuticals, Faeth Therapeutics, Selectin Therapeutics and Senescea Therapeutics. R.P.K. is a cofounder of and consultant for Econic Biosciences. R.C. is a consultant for Sanavia Oncology and LevitasBio. D.P. is on the scientific advisory board of Insitro. J.M. holds equity in Scholar Rock. None of these affiliations represent a conflict of interest with respect to the design or execution of this study or interpretation of data presented in this report. The other authors declare no competing interests.
Figures















References
MeSH terms
Substances
Grants and funding
- T32 CA160001/CA/NCI NIH HHS/United States
- P01CA13106/U.S. Department of Health & Human Services | NIH | National Cancer Institute (NCI)
- F32CA177072/U.S. Department of Health & Human Services | NIH | National Cancer Institute (NCI)
- PF-13-037-01-DMC/American Cancer Society (American Cancer Society, Inc.)
- W81XWH-22-1-0911/U.S. Department of Defense (United States Department of Defense)
- F32 CA177072/CA/NCI NIH HHS/United States
- R35 CA252978/CA/NCI NIH HHS/United States
- LCF/BQ/PI20/11760006/"la Caixa" Foundation (Caixa Foundation)
- P01 CA013106/CA/NCI NIH HHS/United States
- 101041659/Edward P. Evans Foundation
- K99 CA266939/CA/NCI NIH HHS/United States
- T32CA160001/U.S. Department of Health & Human Services | NIH | National Cancer Institute (NCI)
- K99CA266939/U.S. Department of Health & Human Services | NIH | National Cancer Institute (NCI)
- P30 CA008748/CA/NCI NIH HHS/United States
- DRG-2383-19/Damon Runyon Cancer Research Foundation (Cancer Research Fund of the Damon Runyon-Walter Winchell Foundation)
- P30CA08748/U.S. Department of Health & Human Services | NIH | National Cancer Institute (NCI)
- R35CA252978/U.S. Department of Health & Human Services | NIH | National Cancer Institute (NCI)
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

