IGF2BP3 redirects glycolytic flux to promote one-carbon metabolism and RNA methylation
- PMID: 41015030
- PMCID: PMC12593248
- DOI: 10.1016/j.celrep.2025.116330
IGF2BP3 redirects glycolytic flux to promote one-carbon metabolism and RNA methylation
Abstract
Insulin-like growth factor 2 mRNA-binding protein 3 (IGF2BP3), an oncofetal RNA-binding protein and a non-canonical reader of N6-methyladenosine (m6A) mRNA modifications, is known to be critical for leukemogenesis. To understand how the oncogenic function of IGF2BP3 impacts metabolism, we performed metabolic profiling and observed changes in glycolytic flux and one-carbon metabolism, including the biosynthesis of S-adenosyl methionine (SAM), a key substrate for methylation reactions within the cell. Using enhanced crosslinking immunoprecipitation (eCLIP) and polyribosome profiling, we found that IGF2BP3 promotes translation of the regulatory subunit of the methionine adenosyltransferase complex (MAT2B), which is involved in SAM production. Remarkably, IGF2BP3 promotes and alters the level and pattern of m6A modifications on mRNA. Taken together, these data suggest the intriguing hypothesis that IGF2BP3 rewrites the epitranscriptome in leukemia cells. Furthermore, this work highlights an interconnection between oncogenic metabolism and RNA modifications, suggesting that pervasive gene expression changes necessary for oncogenesis may be perpetuated by post-transcriptional gene regulation.
Keywords: B-acute lymphoblastic leukemia; CP: Metabolism; MLL-rearranged leukemia; RNA-binding protein; epitranscriptomics.
Copyright © 2025 The Authors. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests D.S.R. has served as a consultant to AbbVie, a pharmaceutical company that develops and markets drugs for hematologic disorders. D.S.R., N.K.G., G.M.S., J.P.S., R.D.D., M.L.T., and A.K.J. are inventors on a patent application that includes the compound I3IN-002.
Figures
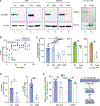





Update of
-
Metabolic regulation of RNA methylation by the m6A-reader IGF2BP3.bioRxiv [Preprint]. 2024 Nov 3:2024.10.31.621399. doi: 10.1101/2024.10.31.621399. bioRxiv. 2024. Update in: Cell Rep. 2025 Oct 28;44(10):116330. doi: 10.1016/j.celrep.2025.116330. PMID: 39554138 Free PMC article. Updated. Preprint.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

