Complete loss of SLC30A8 in humans improves glucose metabolism and beta cell function
- PMID: 41020949
- PMCID: PMC12594714
- DOI: 10.1007/s00125-025-06530-3
Complete loss of SLC30A8 in humans improves glucose metabolism and beta cell function
Abstract
Aims/hypothesis: Genetic association studies have demonstrated that partial loss of SLC30A8 Function protects against type 2 diabetes in humans. We investigated the impact of complete loss of SLC30A8 Function on type 2 diabetes risk and related phenotypes in humans.
Methods: The Pakistan Genome Resource (PGR), a biobank comprising whole-exome and whole-genome sequences of 145,037 participants, was analysed for phenotypic associations with SLC30A8 loss-of-function (LoF) variants. To follow up on the observations in the PGR, we conducted recall-by-genotype analyses of SLC30A8 LoF heterozygotes and homozygotes, as well as their participating family members, using OGTTs.
Results: We identified 18 SLC30A8 knockouts, including homozygotes for a variant enriched in South Asians (Gln174Ter), and 1024 heterozygotes for LoF variants. Type 2 diabetes risk was lower in SLC30A8 LoF heterozygotes and homozygotes relative to non-carriers, and the protective effect strengthens in a gene dose-dependent manner (ORadditive=0.62; 95% CI 0.53, 0.72; p=1.1×10-9; ORrecessive=0.34; 95% CI 0.12, 0.93; p=0.04). OGTTs in recall-by-genotype studies showed a gene dose-dependent reduction in glucose levels, coupled with elevated insulin.
Conclusions/interpretation: The corrected insulin response, disposition index and insulin sensitivity index in LoF heterozygotes and homozygotes indicated higher glucose-stimulated insulin secretion with preserved beta cell function that was independent of BMI. These data suggest that therapeutic inhibition of SLC30A8, up to and including complete knockout, may treat type 2 diabetes safely and effectively.
Keywords: SLC30A8; Diabetes; Knockout; Pakistan Genome Resource; South Asian; ZnT8.
© 2025. The Author(s).
Conflict of interest statement
Acknowledgements: The authors would like to thank V. George, P. Runge and P. Schroeder of Novartis for their contributions to compliance documentation associated with human tissue samples. The authors are additionally grateful to the PGR participants, recall-by-genotype participants, and UK Biobank participants for their vital contributions to this research. Data availability: The whole-exome sequencing data that we have generated includes rare LoF variants, many that have a count of fewer than five, which may potentially lead to identification of study participants. All academic requests to access relevant data should be sent to ks76@cncdpk.com. The Center for Non-Communicable Diseases (CNCD) will ask investigators to sign a data confidentiality agreement that would require investigators to maintain de-identification of the study participants. Funding: DS has received grants from the National Institutes of Health ( www.nih.gov ) (R01-HL-145437, R01-HG-010689, R01-HL133339, X01HL139399, RC2 HL101834-01 and RC1 TW008485-01). DS has also received funding from Regeneron Pharmaceuticals, Eli Lilly & Company, Merck, Astra Zeneca, NGM Biopharmaceuticals Inc., GSK, Astellas Pharma Inc. and Novo Nordisk. Whole-exome sequencing of the Pakistan Genome Resource was performed at Regeneron with support from Regeneron, Novartis, AstraZeneca and Astellas. Authors’ relationships and activities: DS has received funding from Regeneron Pharmaceuticals, Eli Lilly & Company, Novartis, Merck, Astra Zeneca, NGM Biopharmaceuticals Inc., GSK, Astellas Pharma Inc. and Novo Nordisk. AMG, CK, LBL, RZ, MEC, DPD, ABG and JED are employees of Novartis. IS was an employee of Novartis at the time that this study was performed and is currently an employee of Yarrow Biotechnology. AA and JLH are employees of AstraZeneca. CU was an employee of AstraZeneca at the time that the study was performed and is currently an employee of Novo Nordisk. KT, MA and IK are employees of Astellas Pharma. ARS and JLR-F are employees of the Regeneron Genetics Center. The remaining authors declare that there are no relationships or activities that might bias, or be perceived to bias, their work. Contribution statement: LBL, CK, SK, MZK, AMG, JED and DS contributed to the study design, drafting and critical revision of the article; LBL, CK, SK, MZK, RZ, MEC, DPD, ABG, IS, AA, JLH, CU, KT, MA, IK, JLR-F, ARS, AR, MJ, MRM, MBL, UA, RS, AJ, MHS, SA, FRM and MI contributed to data collection and analysis. All authors reviewed the article and approved the final version to be published. DS is the guarantor of this work, and, as such, had full access to all the data in the study and takes responsibility for the integrity of the data and the accuracy of the data analysis.
Figures
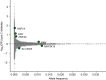
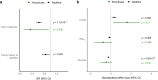

References
-
- Sladek R, Rocheleau G, Rung J et al (2007) A genome-wide association study identifies novel risk loci for type 2 diabetes. Nature 445(7130):881–885. 10.1038/nature05616 - PubMed
-
- Cauchi S, Del Guerra S, Choquet H et al (2010) Meta-analysis and functional effects of the SLC30A8 rs13266634 polymorphism on isolated human pancreatic islets. Mol Genet Metab 100(1):77–82. 10.1016/j.ymgme.2010.01.001 - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Research Materials

