multiSMD - A Python Toolset for Multidirectional Steered Molecular Dynamics
- PMID: 41037437
- PMCID: PMC12570128
- DOI: 10.1021/acs.jcim.5c01742
multiSMD - A Python Toolset for Multidirectional Steered Molecular Dynamics
Abstract
Molecular forces govern all biological processes from cellular mechanics to molecular recognition events. Understanding the direction-dependence of these forces is particularly critical for elucidating fundamental interactions, such as protein-protein binding, ligand dissociation, and signal mechanotransduction. While steered molecular dynamics (SMD) simulations enable the study of force-induced transitions, conventional single-direction approaches may overlook anisotropic mechanical responses inherent to biomolecular systems. Therefore, probing the mechanical stability of molecular systems with respect to a director of an external force may provide critical information. Here, we present multiSMD, a Python-based tool that automates the setup and analysis of multidirectional SMD simulations in NAMD and GROMACS. By systematically probing forces along multiple spatial vectors, multiSMD captures direction-dependent phenomena, such as changing energy barriers or structural resilience, that remain hidden in standard SMD. We demonstrate the utility of our approach through three distinct applications: (i) anisotropic unbinding in a protein-protein complex, (ii) search for ligand dissociation pathways dependent on the pulling direction, and (iii) force-induced remodeling of intrinsically disordered regions in proteins. multiSMD streamlines the exploration of nanomechanical anisotropy in biomolecules, offering a computational framework to guide experiments (e.g., atomic force microscopy - AFM or optical tweezers) and uncover mechanistic properties inaccessible to single-axis methods.
Figures
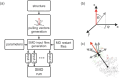

References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

