NAA60 facilitates LRRC8A- and LRRC8D-mediated platinum drug uptake
- PMID: 41053424
- PMCID: PMC12501270
- DOI: 10.1038/s42003-025-08826-x
NAA60 facilitates LRRC8A- and LRRC8D-mediated platinum drug uptake
Abstract
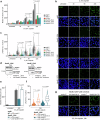
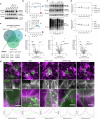
The platinum-based drugs cis- and carboplatin, which are crucial for treating cancers with DNA repair defects, like those caused by BRCA1/2 mutations, rely on the volume-regulated anion channel subunits LRRC8A and LRRC8D for about 50% of cellular drug uptake. Yet, the precise mechanisms of how LRRC8A and LRRC8D mediate this function are largely unknown. Here, we identify NAA60, an N-terminal acetyltransferase, which localizes to the Golgi apparatus to affect LRRC8A and LRRC8D function. Our data suggest that NAA60 acetylates the LRRC8A/D N-termini, and its loss decreases cis- and carboplatin uptake resulting in drug resistance of otherwise hypersensitive BRCA1;p53-deficient cells and tumors. Furthermore, we mimicked the absence of the neutralizing acetyl moiety that is observed after loss of NAA60 by introducing positively charged amino acids at the N-termini of LRRC8A/D, which indeed decreased cis- and carboplatin sensitivity. Our findings highlight the importance of N-terminal acetylation by NAA60 for effective platinum drug uptake, offering new insights into overcoming drug resistance.
© 2025. The Author(s).
Conflict of interest statement
Competing interests: The authors declare no competing interests.
Figures





References
-
- Corvaja, C., Garutti, M., Gerratana, L., Pelizzari, G. & Puglisi, F. Hype or hope? The strange case of platinum salts’ renaissance in breast cancer. Expert Rev. Anticancer Ther.19, 1005–1008 (2019). - PubMed
-
- Postel-Vinay, S. et al. The potential of exploiting DNA-repair defects for optimizing lung cancer treatment. Nat. Rev. Clin. Oncol.9, 144–155 (2012). - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

