A nanobody-based tri-specific NK cell engager targeting CD5 triggers antitumor immunity
- PMID: 41067231
- PMCID: PMC12629802
- DOI: 10.1016/j.xcrm.2025.102409
A nanobody-based tri-specific NK cell engager targeting CD5 triggers antitumor immunity
Abstract
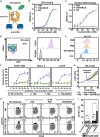
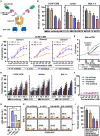
The poor prognosis of patients with recurrent or refractory T cell malignancies emphasizes the need for improved immunotherapies. CD5 is a characteristic marker of malignant T cells and is expressed on almost all normal T cells. Therefore, for treating T cell malignancies, focusing on natural killer (NK) cells lacking CD5 expression may elicit a better safety profile than that by T cell-based therapies. We generate a CD5-targeted NK cell engager (NKCE) through the specific binding of CD16a nanobody and a high-affinity anti-CD5 antibody. Its antitumor potency is demonstrated in vitro. After incorporating interleukin (IL)-15Rα/IL-15, the modified tri-NKCE exhibits stronger antitumor efficacy against CD5+ malignant tumor cells, with the production of more cytokines and chemokines. In vivo, tri-NKCE exhibits stronger cytotoxicity by enhancing NK cell proliferation. Compared with chimeric antigen receptor (CAR)-T cells, this tri-NKCE exhibits no toxicity to normal T cells. In conclusion, tri-NKCE offers a safer and cost-effective immunotherapy against T cell malignancies.
Keywords: CD16a; CD5; T cell malignancies; immunotherapy; nanobody; natural killer cells.
Copyright © 2025 The Authors. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests The authors declare no competing interests.
Figures






References
-
- Einsiedel H.G., von Stackelberg A., Hartmann R., Fengler R., Schrappe M., Janka-Schaub G., Mann G., Hählen K., Göbel U., Klingebiel T., et al. Long-term outcome in children with relapsed ALL by risk-stratified salvage therapy: results of trial acute lymphoblastic leukemia-relapse study of the Berlin-Frankfurt-Münster Group 87. J. Clin. Oncol. 2005;23:7942–7950. doi: 10.1200/JCO.2005.01.1031. - DOI - PubMed
-
- Winter S.S., Dunsmore K.P., Devidas M., Wood B.L., Esiashvili N., Chen Z., Eisenberg N., Briegel N., Hayashi R.J., Gastier-Foster J.M., et al. Improved survival for children and young adults with T-lineage acute lymphoblastic leukemia: results from the Children’s Oncology Group AALL0434 Methotrexate Randomization. J. Clin. Oncol. 2018;36:2926–2934. doi: 10.1200/JCO.2018.77.7250. - DOI - PMC - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

