VRK2 targeting potentiates anti-PD-1 immunotherapy in hepatocellular carcinoma through MYC destabilization
- PMID: 41073389
- PMCID: PMC12514316
- DOI: 10.1038/s41467-025-64079-6
VRK2 targeting potentiates anti-PD-1 immunotherapy in hepatocellular carcinoma through MYC destabilization
Abstract
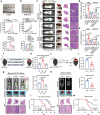
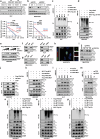
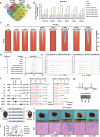
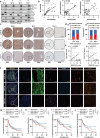
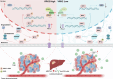
Dysregulation of MYC proto-oncogene, bHLH transcription factor (MYC) represents a common yet mechanistically unresolved driver of hepatocellular carcinoma (HCC). While MYC remains an elusive therapeutic target, developing strategies to promote its degradation emerges as a promising alternative approach. Here we show that vaccinia-related kinase 2 (VRK2) functions as a direct MYC-interacting kinase that stabilizes the oncoprotein through phosphorylation at Serine (Ser)281/293. This phosphorylation enables VRK2 to compete with the Skp1-Cullin-F-box protein complex containing FBXO24 (SCF-FBXO24) E3 ligase, thereby blocking MYC polyubiquitination and proteasomal degradation. The stabilized MYC-VRK2 complex amplifies transcriptional activation of protumorigenic programs, including the immune checkpoint programmed cell death ligand 1 (PD-L1) and VRK2 itself, establishing a self-reinforcing oncogenic circuit. Therapeutic inhibition of VRK2 in HCC models reduces MYC protein levels, suppresses tumor progression, and synergizes with anti- programmed cell death-1 (PD-1) immunotherapy. Our results reveal VRK2-mediated stabilization of MYC as a critical nexus linking hepatocarcinogenesis to immune evasion, proposing VRK2 kinase inhibition as a mechanism-based therapeutic strategy for MYC-driven HCC.
© 2025. The Author(s).
Conflict of interest statement
Competing interests: The authors declare no competing interests.
Figures




References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials

