Cardiomyocyte lncRNA Cpat maintains cardiac homeostasis and mitochondria function by targeting citrate synthase acetylation
- PMID: 41073440
- PMCID: PMC12513993
- DOI: 10.1038/s41467-025-64072-z
Cardiomyocyte lncRNA Cpat maintains cardiac homeostasis and mitochondria function by targeting citrate synthase acetylation
Abstract
Myocardial energy metabolism disorders are essential pathophysiology in sepsis-associated myocardial injury. Yet, the underlying mechanisms involving impaired mitochondrial respiratory function upon myocardial injury remain poorly understood. Here we identify an unannotated and cardiomyocyte-enriched long non-coding RNA, Cpat (cardiac-protector-associated transcript), that plays an important role in regulating the dynamics of cardiomyocyte mitochondrial tricarboxylic acid (TCA) cycle. Cpat is essential to the mitochondrial respiratory function by targeting key metabolic enzymes and modulating TCA cycle flux. Specifically, Cpat enhances the association of TCA cycle core components malate dehydrogenase (MDH2), citrate synthase (CS), and aconitase (ACO2). Acetyltransferase general control non-repressed protein-5 (GCN5) acetylates CS and destabilizes the MDH2-CS-ACO2 complex formation. Cpat inhibits this GCN5 activity and facilitates MDH2-CS-ACO2 complex formation and TCA cycle flux. We reveal that Cpat-mediated mitochondrial metabolic homeostasis is vital in mitigating myocardial injury in sepsis-induced cardiomyopathy, positioning Cpat as a promising therapeutic target for preserving myocardial cellular metabolism and function.
© 2025. The Author(s).
Conflict of interest statement
Competing interests: The authors declare no competing interests.
Figures

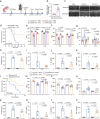





References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

