Microtubule forces drive nuclear damage in LMNA cardiomyopathy
- PMID: 41073815
- PMCID: PMC12611788
- DOI: 10.1038/s44161-025-00727-w
Microtubule forces drive nuclear damage in LMNA cardiomyopathy
Abstract
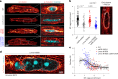
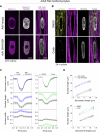
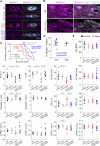
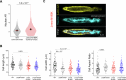
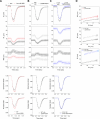
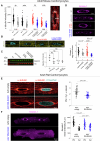
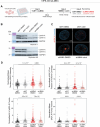
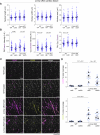
Nuclear homeostasis requires balanced forces between the cytoskeleton and the nucleus. Mutations in LMNA, which encodes lamin A/C, weaken the nuclear lamina, leading to nuclear damage and muscle disease. Disrupting the linker of nucleoskeleton and cytoskeleton (LINC) complex, which connects the cytoskeleton to the nucleus, may ameliorate LMNA-associated cardiomyopathy, yet the cardioprotective mechanism remains unclear. Here we developed an assay to quantify the coupling between cardiomyocyte contraction and nuclear deformation and interrogate its dependence on the nuclear lamina and LINC complex. The LINC complex was mostly dispensable for transferring contractile strain to the nucleus, and its disruption did not rescue elevated nuclear strain in lamin A/C-deficient cardiomyocytes. Instead, LINC complex disruption eliminated the microtubule cage encircling the nucleus. Microtubule disruption prevented nuclear damage and preserved cardiac function in lamin A/C deficiency. Computational modeling revealed that microtubule forces create local stress concentrations that damage lamin A/C-deficient nuclei. These findings identify microtubule-dependent force transmission as a pathological driver and therapeutic target for LMNA cardiomyopathy.
© 2025. The Author(s).
Conflict of interest statement
Competing interests: The authors declare no competing interests.
Figures









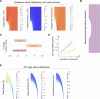
Update of
-
Microtubule forces drive nuclear damage in LMNA cardiomyopathy.bioRxiv [Preprint]. 2025 Jun 2:2024.02.10.579774. doi: 10.1101/2024.02.10.579774. bioRxiv. 2025. Update in: Nat Cardiovasc Res. 2025 Nov;4(11):1501-1520. doi: 10.1038/s44161-025-00727-w. PMID: 38948795 Free PMC article. Updated. Preprint.
References
-
- Zhou, C., Rao, L., Shanahan, C. M. & Zhang, Q. Nesprin-1/2: roles in nuclear envelope organisation, myogenesis and muscle disease. Biochem. Soc. Trans.46, 311–320 (2018). - PubMed
-
- Ross, J. A. & Stroud, M. J. The nucleus: mechanosensing in cardiac disease. Int. J. Biochem. Cell Biol.137, 106035 (2021). - PubMed
MeSH terms
Substances
Grants and funding
- R01 HL149891/HL/NHLBI NIH HHS/United States
- URoL-2022048/National Science Foundation (NSF)
- HL149891/U.S. Department of Health & Human Services | NIH | National Heart, Lung, and Blood Institute (NHLBI)
- GM153257/U.S. Department of Health & Human Services | NIH | National Institute of General Medical Sciences (NIGMS)
- 20CVD01/Fondation Leducq
- HL133080/U.S. Department of Health & Human Services | NIH | National Heart, Lung, and Blood Institute (NHLBI)
- 15-48571/National Science Foundation (NSF)
- R01 HL082792/HL/NHLBI NIH HHS/United States
- R01 HL133080/HL/NHLBI NIH HHS/United States
- R01 AR084664/AR/NIAMS NIH HHS/United States
- 1715606/NSF | ENG/OAD | Division of Chemical, Bioengineering, Environmental, and Transport Systems (CBET)
- R35 GM153257/GM/NIGMS NIH HHS/United States
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous
