Engineered 3D immuno-glial-neurovascular human miBrain model
- PMID: 41105712
- PMCID: PMC12557797
- DOI: 10.1073/pnas.2511596122
Engineered 3D immuno-glial-neurovascular human miBrain model
Abstract
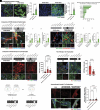
Patient-specific, human-based cellular models integrating a biomimetic blood-brain barrier, immune, and myelinated neuron components are critically needed to enable accelerated, translationally relevant discovery of neurological disease mechanisms and interventions. To construct a human cell-based model that includes these features and all six major brain cell types needed to mimic disease and dissect pathological mechanisms, we have constructed, characterized, and utilized a multicellular integrated brain (miBrain) immuno-glial-neurovascular model by engineering a brain-inspired 3D hydrogel and identifying conditions to coculture these six brain cell types, all differentiated from patient induced pluripotent stem cells. miBrains recapitulate in vivo-like hallmarks inclusive of neuronal activity, functional connectivity, barrier function, myelin-producing oligodendrocyte engagement with neurons, multicellular interactions, and transcriptomic profiles. We implemented the model to study Alzheimer's Disease pathologies associated with APOE4 genetic risk. APOE4 miBrains differentially exhibit amyloid aggregation, tau phosphorylation, and astrocytic glial fibrillary acidic protein. Unlike the coemergent fate specification of glia and neurons in other organoid approaches, miBrains integrate independently differentiated cell types, a feature we harnessed to identify that APOE4 in astrocytes promotes neuronal tau pathogenesis and dysregulation through crosstalk with microglia.
Keywords: biomaterials; brain organoid; microphysiological system; neuro-immune; neurovascular.
Conflict of interest statement
Competing interests statement:The authors A.E.S, A.B., J.W.B., R.L., and L.-H.T. have filed a patent application (PCT/US2021/047853) based on the findings.
Figures



Update of
-
Engineered 3D Immuno-Glial-Neurovascular Human miBrain Model.bioRxiv [Preprint]. 2024 Jun 11:2023.08.15.553453. doi: 10.1101/2023.08.15.553453. bioRxiv. 2024. Update in: Proc Natl Acad Sci U S A. 2025 Oct 21;122(42):e2511596122. doi: 10.1073/pnas.2511596122. PMID: 37645757 Free PMC article. Updated. Preprint.
References
MeSH terms
Grants and funding
- 1F32AG072813-01/HHS | NIH (NIH)
- P30-CA14051/HHS | NIH | National Cancer Institute (NCI)
- UG3 NS115064/NS/NINDS NIH HHS/United States
- 3-UG3-NS115064-01S1/HHS | NIH (NIH)
- K01 AG083734/AG/NIA NIH HHS/United States
- BT Charitable Foundation/BT Charitable Foundation
- Picower Fellowship/Picower Fellowship
- 2176252/Kavanaugh Fellowship
- F32 AG072813/AG/NIA NIH HHS/United States
- Schmidt Science Fellows in partnership with the Rhodes Trust/Schmidt Science Fellows in partnership with the Rhodes Trust
- 1K01AG083734-01/HHS | NIH (NIH)
- P30 CA014051/CA/NCI NIH HHS/United States
- UH3 NS115064/NS/NINDS NIH HHS/United States
LinkOut - more resources
Full Text Sources

