Chelerythrine inhibits esophageal squamous cell carcinoma progression via PINK1-Parkin-mediated mitophagy
- PMID: 41107946
- PMCID: PMC12532471
- DOI: 10.1186/s12967-025-07025-w
Chelerythrine inhibits esophageal squamous cell carcinoma progression via PINK1-Parkin-mediated mitophagy
Abstract
Background: Chelerythrine (CHE) exhibits notable anti-inflammatory and antitumor properties, while its impact on esophageal squamous cell carcinoma (ESCC), especially the underlying mechanisms remain unclear. In this study, we aim to investigate the roles and mechanism of CHE in the ESCC treatment.
Methods: Human ESCC cell lines and organoids were used for in vitro cell experiments, BALB/c nude mice were used for in vivo animal experiments. To investigate the underlying mechanism of CHE treatment, drug library screen, RNA sequencing analysis, TMT-based quantitative proteomic analysis, western blotting analysis, immunofluorescence, immunohistochemistry, quantitative real-time polymerase chain reaction, mitochondrial membrane potential assay, apoptosis assay, detection of mitochondrial reactive oxygen species (mtROS), autophagic flux monitoring, transmission electron microscopy, and seahorse XF-96 metabolic flux analysis were used to assess the effect of CHE and relevant mechanism.
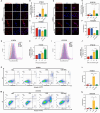
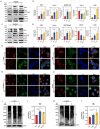
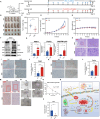
Results: CHE dose-dependently inhibited the proliferation, migration, and invasion of ESCC cells. CHE also induced cell apoptosis and triggered PTEN-induced kinase 1 (PINK1)-Parkin-mediated mitophagy-mediated cell death by elevating the production of reactive oxygen species in mitochondria and diminishing mitochondrial membrane potential (MMP). However, the production of autophagosomes and autolysosomes induced by CHE altered when used in combination with the autophagy inhibitors 3-methyladenine (3-MA) or bafilomycin A1 (BafA1), indicating that it induced complete autophagic flux in the cells. Mechanistically, CHE affected multiple signaling pathways associated with ubiquitin-mediated proteolysis, mitophagy, and mitochondrial energy metabolism, indicating its close involvement in mitophagy occurrence. In addition, CHE treatment significantly reduced tumor size and weight in nude mice bearing KYSE150 tumors and retarded the growth of organoids derived from patients, it also reduced the ratio of M2 macrophage in tumor microenvironment and cell metabolism.
Conclusions: CHE activates PINK1-Parkin-mediated mitophagy and disrupts mitochondrial homeostasis, and it also affects the tumor environment and cell metabolism, ultimately leading to cell death, supporting the potential of CHE for ESCC therapy.
Keywords: Chelerythrine; Esophageal squamous cell carcinoma; Mitophagy; PINK1-Parkin pathway.
© 2025. The Author(s).
Conflict of interest statement
Declarations. Ethics approval and consent to participate: All mouse experiments were performed in compliance with the ethical policies and procedures approved by the Institutional Animal Care and Use Committee of Xiamen University (Approval no. XUMLAC20200202). Animal experiment procedures were performed according to the Guide for the Care and Use of Laboratory Animals of the National Institutes of Health and followed the guidelines of the Animal Welfare Act. Consent for publication: All authors have read and approved the final manuscript. Competing interests: The authors declare that they have no competing interests.
Figures





References
-
- Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, Bray F. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2021;71:209–49. - PubMed
-
- Wu J, Wang L, Du X, Sun Q, Wang Y, Li M, Zang W, Liu K, Zhao G. alpha-solanine enhances the chemosensitivity of esophageal cancer cells by inducing microRNA–138 expression. Oncol Rep. 2018;39:1163–72. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials

