Genome-wide association study of childhood B-cell acute lymphoblastic leukemia reveals novel African ancestry-specific susceptibility loci
- PMID: 41125582
- PMCID: PMC12546916
- DOI: 10.1038/s41467-025-64337-7
Genome-wide association study of childhood B-cell acute lymphoblastic leukemia reveals novel African ancestry-specific susceptibility loci
Abstract
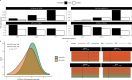
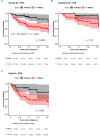
B-cell acute lymphoblastic leukemia (B-ALL) is the most common pediatric malignancy. Given racial/ethnic differences in incidence and outcomes, B-ALL genome-wide association studies among children of African ancestry are needed. Leveraging multi-institutional datasets with 840 African American children with B-ALL and 3360 controls, nine loci achieved genome-wide significance (P < 5 × 10-8) after meta-analysis. Two loci were established trans-ancestral susceptibility regions (IKZF1, ARID5B), while the remaining novel loci were specific to African populations. Five-year overall survival among children carrying novel risk alleles was significantly worse (83% versus 96% in non-carriers, P = 4.8 × 10-3). Novel risk variants were also associated with subtype-specific disease (P < 0.05), including higher susceptibility for a subtype overrepresented in African American children (TCF3-PBX1) and lower susceptibility for a subtype with excellent prognosis (ETV6-RUNX1). Functional experiments revealed novel B-ALL risk variants had allele-specific differences in transcriptional activity (P < 0.05) in B-cell and leukemia cell lines. These findings shed insights into ancestry-related differences in leukemogenesis and prognosis.
© 2025. The Author(s).
Conflict of interest statement
Competing interests: The authors declare no competing interests.
Figures



References
-
- Howlader, N. et al. SEER cancer statistics review, 1975–2018. National Cancer Institute (2021).
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

