Probing the core metabolism of Cereibacter sphaeroides by transposon mutagenesis
- PMID: 41165417
- PMCID: PMC12632250
- DOI: 10.1128/jb.00306-25
Probing the core metabolism of Cereibacter sphaeroides by transposon mutagenesis
Abstract
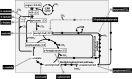
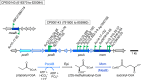
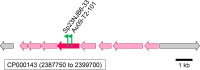
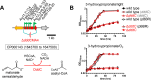
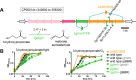
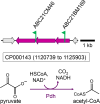
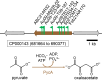
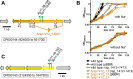
During phototrophic growth, Cereibacter sphaeroides can use several carbon substrates that are central carbon intermediates (e.g., succinate and L-malate) or that require only a few steps to enter central carbon metabolism (e.g., acetate and D-malate). In addition, with light as the energy source, the carbon substrate provided will function as a carbon source for cell carbon synthesis only. Therefore, C. sphaeroides is ideally suited to understand the changes necessary to switch between different carbon sources and, consequently, to redirect carbon flow in central carbon metabolism. This study describes C. sphaeroides transposon mutants that have lost the ability to use one or more of the organic carbon sources 3-hydroxypropionate, acetate, L-malate, propionate/HCO3-, butyrate/HCO3, L-lactate, D-lactate, D-malate, and L-glutamate. Pyruvate carboxylase and pyruvate dehydrogenase were confirmed to connect the precursor metabolite pools of pyruvate and oxaloacetate or acetyl-CoA, respectively, as was the ethylmalonyl-CoA pathway connecting acetyl-CoA and oxaloacetate pools. Transposon and in-frame deletion mutants suggest that 3-hydroxypropionate is oxidized to CO2 and acetyl-CoA, involving a malonate semialdehyde dehydrogenase. The presence of this oxidative route makes pyruvate dehydrogenase dispensable during 3-hydroxypropionatedependent growth. Therefore, acetyl-CoA represents a second entry point into central carbon metabolism for 3-hydroxypropionate besides succinyl-CoA, and it is proposed that the simultaneous functioning of the two routes minimizes transiently produced CO2/HCO3-. Another significant outcome of this study is the identification of genes encoding a L-glutamate TRipartite ATP-independent transporter, which was characterized biochemically 30 years ago.IMPORTANCESeveral aspects of the process of carbon assimilation, defined as the conversion of a carbon source into cell carbon, are conserved throughout life. For example, common building blocks give rise to proteins and nucleic acids, and the carbon for building blocks, cofactors, and secondary metabolites is derived from common precursor metabolites such as acetyl-CoA, pyruvate, or oxaloacetate. Using carbon substrates that require only one or a few steps to enter central carbon metabolism facilitates insights into the changes that occur to accommodate growth with different carbon substrates. In this study, transposon mutants that affect carbon flow in the core metabolism of Cereibacter sphaeroides were identified. Apparent redundancies of pathways can be explained by the need to maintain overall redox balance.
Keywords: 3-hydroxypropionate metabolism; central carbon metabolism; pyruvate carboxylase; pyruvate dehydrogenase; secondary L-glutamate TAXI transport system.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
References
-
- Hördt A, López MG, Meier-Kolthoff JP, Schleuning M, Weinhold L-M, Tindall BJ, Gronow S, Kyrpides NC, Woyke T, Göker M. 2020. Analysis of 1,000+ type-strain genomes substantially improves taxonomic classification of Alphaproteobacteria. Front Microbiol 11:468. doi: 10.3389/fmicb.2020.00468 - DOI - PMC - PubMed
-
- Molisch H. 1907. Die Purpurbakterien : nach neuen Untersuchungen; eine mikrobiologische Studie. Verlag von Gustav Fischer, Jena.
-
- Muller FM. 1933. On the metabolism of the purple sulphur bacteria in organic media. Archiv Mikrobiol 4:131–166. doi: 10.1007/BF00407535 - DOI
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

