Data navigation on the ENCODE portal
- PMID: 41168159
- PMCID: PMC12575607
- DOI: 10.1038/s41467-025-64343-9
Data navigation on the ENCODE portal
Abstract
Spanning two decades, the collaborative ENCODE project aims to identify all the functional elements within human and mouse genomes. To best serve the scientific community, the comprehensive ENCODE data including results from 23,000+ functional genomics experiments, 800+ functional elements characterization experiments and 60,000+ results from integrative computational analyses are available on an open-access data-portal ( https://www.encodeproject.org/ ). The final phase of the project includes data from several novel assays aimed at characterization and validation of genomic elements. In addition to developing and maintaining the data portal, the Data Coordination Center (DCC) implemented and utilised uniform processing pipelines to generate uniformly processed data. Here we report recent updates to the data portal including a redesigned home page, an improved search interface, new custom-designed pages highlighting biologically related datasets and an enhanced cart interface for data visualisation plus user-friendly data download options. A summary of data generated using uniform processing pipelines is also provided.
© 2025. The Author(s).
Conflict of interest statement
Competing interests: The authors have no competing interests.
Figures



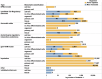



References
-
- The modENCODE Project. Genome.govhttps://www.genome.gov/26524507/the-modencode-project-model-organism-enc....
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

