Tissue tropism and functional adaptation of the SARS-CoV-2 spike protein in a fatal case of COVID-19
- PMID: 41170974
- PMCID: PMC12645954
- DOI: 10.1128/jvi.00857-25
Tissue tropism and functional adaptation of the SARS-CoV-2 spike protein in a fatal case of COVID-19
Abstract
Systemic spread of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) to extrapulmonary tissues has been observed following acute infections. Autopsy studies further indicate tissue-specific virus diversity, including in immune-privileged sites. Questions remain on the viral dynamics leading to the tissue tropism of SARS-CoV-2, including evolutionary trajectories and functional adaptations that could impact persistence and transmission. In this study, we characterized SARS-CoV-2 genomes from 27 distinct tissues collected from an autopsy case where the patient had a primary immune deficiency. We identified tissue-specific virus genotypes, in some instances coexisting within the same sites, with mutations primarily in the receptor-binding domain of the spike protein. Protein simulations and isolation of infectious virus indicate combinations of spike substitutions that would lead to increased protein stability and stronger binding of the virus to host cells. This highlights the importance of studying patients with weakened immune responses where potential tissue reservoirs provide an environment permissive for SARS-CoV-2 evolution and diversification.IMPORTANCEPersistent severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) infections in immunocompromised individuals are considered a potential source of new viral variants. Beyond the respiratory tract, the virus can spread within days to organs like the brain, heart, and kidneys, where distinct tissue microenvironments may further drive viral evolution and the emergence of new mutations. In this study, we compared the genetic diversity of SARS-CoV-2 genomic RNA isolated from 27 distinct tissue sites collected from an individual with a weakened immune system. By linking viral population dynamics across these tissue sites, we defined the extent of compartmentalization during multi-organ spread, highlighting how non-respiratory tissues can impact SARS-CoV-2 diversification.
Keywords: coronavirus; tissue tropism; viral evolution; viral intrahost diversity.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



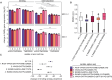

References
-
- Van Cleemput J, van Snippenberg W, Lambrechts L, Dendooven A, D’Onofrio V, Couck L, Trypsteen W, Vanrusselt J, Theuns S, Vereecke N, van den Bosch TPP, Lammens M, Driessen A, Achten R, Bracke KR, Van den Broeck W, Von der Thüsen J, Nauwynck H, Van Dorpe J, Gerlo S, Maes P, Cox J, Vandekerckhove L. 2021. Organ-specific genome diversity of replication-competent SARS-CoV-2. Nat Commun 12:6612. doi: 10.1038/s41467-021-26884-7 - DOI - PMC - PubMed
-
- Normandin E, Rudy M, Barkas N, Schaffner SF, Levine Z, Padera RF Jr, Babadi M, Mukerji SS, Park DJ, MacInnis BL, Siddle KJ, Sabeti PC, Solomon IH. 2023. High-depth sequencing characterization of viral dynamics across tissues in fatal COVID-19 reveals compartmentalized infection. Nat Commun 14:574. doi: 10.1038/s41467-022-34256-y - DOI - PMC - PubMed
-
- Hoffmann M, Kleine-Weber H, Schroeder S, Krüger N, Herrler T, Erichsen S, Schiergens TS, Herrler G, Wu N-H, Nitsche A, Müller MA, Drosten C, Pöhlmann S. 2020. SARS-CoV-2 cell entry depends on ACE2 and TMPRSS2 and is blocked by a clinically proven protease inhibitor. Cell 181:271–280. doi: 10.1016/j.cell.2020.02.052 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

