This is a preprint.
ESX-5 Deletions in Mycobacterium tuberculosis Alter Macrophage Cytokine Signaling and Bacterial Heavy Metal Response
- PMID: 41256704
- PMCID: PMC12621936
- DOI: 10.1101/2025.09.30.679525
ESX-5 Deletions in Mycobacterium tuberculosis Alter Macrophage Cytokine Signaling and Bacterial Heavy Metal Response
Abstract
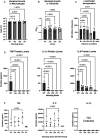
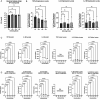
The ESX-5 secretion system is critical for Mycobacterium tuberculosis (Mtb) viability and putatively linked to pathogenesis, but a functional understanding of how it interacts with the host is unknown. ESX-5 is encoded at a single genomic locus with small, paralogous, secreted targets (ESX-5a, 5b, 5c) spaced throughout the genome. To examine host-pathogen interactions of these putative virulence clusters, we made mutant strains lacking these loci and infected primary human macrophages. Surprisingly, all deletion mutants independently reduced cytokine secretion during infection, specific to certain analytes. This defect depended on viable bacteria and was mediated by a post-transcriptional mechanism. In bacterial transcriptomic analyses, each mutant downregulated heavy metal response genes compared to wild type bacteria. Treatment of Mtb with Cu or Cd led to increased ESX-5a and ESX-5c expression, concurrent with increased TNF and IL-6 secretion in macrophages compared to untreated bacilli, indicating a link between ESX-5 expression and cellular cytokine levels.
Conflict of interest statement
Competing Interests Authors have no competing or conflicting interests to declare.
Figures
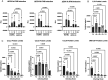
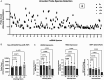

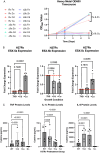
References
-
- Gagneux S. Ecology and evolution of Mycobacterium tuberculosis. Nat Rev Microbiol. 2018. Apr;16(4):202–13. - PubMed
-
- Pai M, Behr MA, Dowdy D, Dheda K, Divangahi M, Boehme CC, et al. Tuberculosis. Nat Rev Dis Primer. 2016. Dec 22;2(1):16076.
-
- Goletti D, Meintjes G, Andrade BB, Zumla A, Lee SS. Insights from the 2024 WHO Global Tuberculosis Report – More Comprehensive Action, Innovation, and Investments required for achieving WHO End TB goals. Int J Infect Dis [Internet]. 2025. Jan 1 [cited 2025 May 1];150. Available from: https://www.ijidonline.com/article/S1201-9712%2824%2900400-4/fulltext
-
- Glaziou P, Sismanidis C, Floyd K, Raviglione M. Global Epidemiology of Tuberculosis. Cold Spring Harb Perspect Med. 2015. Feb;5(2):a017798.
-
- Global Tuberculosis Report 2024. 1st ed. Geneva: World Health Organization; 2024. 1 p.
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
