Structure and expression of a polyubiquitin gene from the crustacean Artemia
- PMID: 7841785
- PMCID: PMC6134374
Structure and expression of a polyubiquitin gene from the crustacean Artemia
Abstract
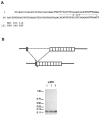
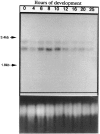
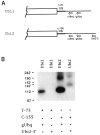
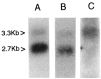
We have characterized two polyubiquitin genes from the crustacean Artemia franciscana. One of them, Ubi1, has nine ubiquitin units and an intron of a minimum size of 3.5 kb that ends 7 bp before the initiator ATG. The 5' end of the transcript from this gene has been identified by anchored PCR. The existence of the other gene (Ubi2) was inferred from several cDNA clones that differ from Ubi1 in the C-terminal extension and in the 3' untranslated region as well as in the nucleotide sequence of the coding region. We find two transcripts of ubiquitin genes, of 2.7 and 3.3 kb. Hybridization of RNA blots with an oligonucleotide specific for Ubi2 gene demonstrates that this gene codes for the 3.3 kb transcript. Ubiquitin messenger RNAs are present in the dormant embryos and their steady-state levels are maximum at 8 h after resumption of development, declining thereafter. The Ubi2 gene transcripts are less abundant but its proportion in relation to the other transcript does not vary with development.
Figures


References
-
- Browne R. A., Sorgeloos P., and Trotman C. N. A. (1991), Artemia Biology, CRC Press, Boca Raton, FL.
-
- Busch H. and Goldknopf L. L. (1981), Mol Cell Biochem 40, 173–187. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
LinkOut - more resources
Full Text Sources
