Analysis of small nuclear RNAs in a precatalytic spliceosome
- PMID: 8836738
- PMCID: PMC6138023
Analysis of small nuclear RNAs in a precatalytic spliceosome
Abstract
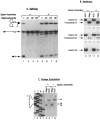
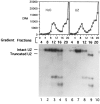
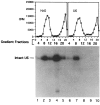
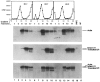
U1 small nuclear RNA plays an important role in early stages of intron recognition and spliceosome assembly. The 5' splice site of the premessenger RNA base-pairs with the 5' end of U1; however, that interaction appears to be replaced by U5 and U6 at later stages of the splicing process. It has not been established when this transition occurs nor what factors are required for the transition. The PRP2 gene of Saccharomyces cerevisiae encodes an RNA-dependent ATPase that belongs to the DEAH putative RNA helicase family. A spliceosome can be assembled in the absence of PRP2 but the ATPase activity of PRP2 is required for the onset of the catalytic reactions in the spliceosome. By probing the precatalytic spliceosome formed in temperature-sensitive prp2 mutant extracts with oligonucleotides complementary to snRNAs, we found that the 5' end of U1 was sensitive to RNase H digestion whereas the 5' splice site-interacting region of U6 became resistant. Furthermore, by treating with heparin, a spliceosome depleted of U1 snRNA was isolated that subsequently underwent splicing with additional protein factors and ATP. Thus, these results indicate that PRP2 is not responsible for the transition from U1 to U6 in the spliceosome and that the entire U1 snRNA is not involved in the catalytic phase of pre-mRNA splicing.
Figures
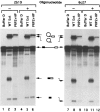
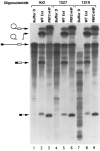
References
-
- Abelson J. RNA processing and the intervening sequence problem. Annu Rev. Biochem. 48:1035–1069; 1979. - PubMed
-
- Brody E.; Abelson J. The spliceosome: Yeast pre-messenger RNA associates with a 40S complex in a splicing-dependent reaction. Science 228:963–967; 1985. - PubMed
-
- Cech T. R. The generality of self-splicing RNA: Relationship to nuclear mRNA splicing. Cell 44:207–210; 1986. - PubMed
-
- Cheng S. C.; Abelson J. Spliceosome assembly in yeast. Genes Dev. 1:1014–1027; 1987. - PubMed
-
- Company M.; Arenas J.; Abelson J. Requirement of the RNA helicase-like protein PRP22 for release of messenger RNA from spliceosomes. Nature 349:487–493; 1991. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
