The essential yeast RNA binding protein Np13p is methylated
- PMID: 8942987
- PMCID: PMC19378
- DOI: 10.1073/pnas.93.24.13641
The essential yeast RNA binding protein Np13p is methylated
Abstract
Arginine methylation is a prevalent modification found in many RNA binding proteins, yet little is known about its functional consequences. Using a monoclonal antibody, 1E4, we have shown that the yeast NPL3 gene product Np13p, an essential RNA binding protein with repeated RGG motifs, is arginine-methylated in vivo. The 1E4 epitope can be generated by incubating recombinant Np13p with partially purified bovine arginine methyltransferase block this reaction. Np13p methylation requires S-adenosyl-L-methionine and also occurs in yeast extracts. An Np13p deletion mutant lacking the RGG domain is not a substrate for methylation, suggesting that the methylation sites lie within the RGG motifs. The discovery of arginine methylation in a genetically tractable organism provides a powerful entrée to understanding the function of this modification, particularly in view of the many roles postulated for Np13p in RNA processing and transport. The recent discovery of phosphorylated serine residues within the RGG domain suggests a hypothesis in which a molecular switch governed by methylation and phosphorylation regulates the biochemical properties of the Np13p RGG domain.
Figures




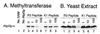

Similar articles
-
Conserved SR protein kinase functions in nuclear import and its action is counteracted by arginine methylation in Saccharomyces cerevisiae.J Cell Biol. 2000 Aug 21;150(4):707-18. doi: 10.1083/jcb.150.4.707. J Cell Biol. 2000. PMID: 10952997 Free PMC article.
-
Novel RING finger proteins, Air1p and Air2p, interact with Hmt1p and inhibit the arginine methylation of Npl3p.J Biol Chem. 2000 Oct 20;275(42):32793-9. doi: 10.1074/jbc.M004560200. J Biol Chem. 2000. PMID: 10896665
-
Arginine methylation of yeast mRNA-binding protein Npl3 directly affects its function, nuclear export, and intranuclear protein interactions.J Biol Chem. 2005 Sep 2;280(35):30888-98. doi: 10.1074/jbc.M505831200. Epub 2005 Jul 5. J Biol Chem. 2005. PMID: 15998636
-
Recent advances in protein methylation: enzymatic methylation of nucleic acid binding proteins.Amino Acids. 1998;15(4):291-306. doi: 10.1007/BF01320895. Amino Acids. 1998. PMID: 9891755 Review.
-
RGG motif proteins: modulators of mRNA functional states.Cell Cycle. 2012 Jul 15;11(14):2594-9. doi: 10.4161/cc.20716. Epub 2012 Jul 15. Cell Cycle. 2012. PMID: 22767211 Free PMC article. Review.
Cited by
-
SRSF1 acts as an IFN-I-regulated cellular dependency factor decisively affecting HIV-1 post-integration steps.Front Immunol. 2022 Nov 15;13:935800. doi: 10.3389/fimmu.2022.935800. eCollection 2022. Front Immunol. 2022. PMID: 36458014 Free PMC article.
-
Nucleocytoplasmic transport of macromolecules.Microbiol Mol Biol Rev. 1997 Jun;61(2):193-211. doi: 10.1128/mmbr.61.2.193-211.1997. Microbiol Mol Biol Rev. 1997. PMID: 9184010 Free PMC article. Review.
-
Conserved SR protein kinase functions in nuclear import and its action is counteracted by arginine methylation in Saccharomyces cerevisiae.J Cell Biol. 2000 Aug 21;150(4):707-18. doi: 10.1083/jcb.150.4.707. J Cell Biol. 2000. PMID: 10952997 Free PMC article.
-
A comprehensive biochemical and genetic analysis of the yeast U1 snRNP reveals five novel proteins.RNA. 1998 Apr;4(4):374-93. RNA. 1998. PMID: 9630245 Free PMC article.
-
Exonic splicing enhancers in fission yeast: functional conservation demonstrates an early evolutionary origin.Genes Dev. 2005 Jan 15;19(2):242-54. doi: 10.1101/gad.1265905. Epub 2004 Dec 29. Genes Dev. 2005. PMID: 15625190 Free PMC article.
References
-
- Clarke S. Curr Opin Cell Biol. 1993;5:977–983. - PubMed
-
- Paik W K, Kim S. Protein Methylation. Boca Raton, FL: CRC; 1990.
-
- Ghosh S K, Paik W K, Kim S. J Biol Chem. 1988;263:19024–19033. - PubMed
-
- Kim S, Chanderkar L P, Ghosh S K. In: Protein Methylation. Paik W K, Kim S, editors. Boca Raton, FL: CRC; 1990. pp. 77–95.
-
- Lischwe M A. In: Protein Methylation. Paik W K, Kim S, editors. Boca Raton, FL: CRC; 1990. pp. 97–123.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

