Identification of a SPH element in the distal region of a human U6 small nuclear RNA gene promoter and characterization of the SPH binding factor in HeLa cell extracts
- PMID: 8979085
- PMCID: PMC6148301
Identification of a SPH element in the distal region of a human U6 small nuclear RNA gene promoter and characterization of the SPH binding factor in HeLa cell extracts
Abstract
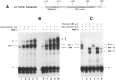
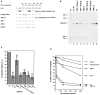
Vertebrate small nuclear RNA (snRNA) gene promoters contain a distal, enhancer-like region that is composed of an octamer motif adjacent to at least one other element. Here we show that a human U6 snRNA distal region contains a SPH motif previously found in several chicken snRNA gene enhancers and the 5'-flanking region of vertebrate selenocysteine tRNA genes. SPH binding factor (SBF) was detected in either chicken or HeLa cell extracts that could bind SPH elements in a species-independent manner. Both human and chicken SBF required divalent cation to bind effectively to DNA. DNase I footprinting experiments indicated that human SBF specifically protected the human U6 SPH element. Furthermore, a SBF polypeptide of approximately 85 kDa was detected in both HeLa and chicken extracts following ultraviolet light-mediated cross-linking to human U6 or chicken U4 SPH elements. A part of the human U6 SPH element was quite sensitive to mutation, as demonstrated by both specific protein binding and transcription assays. From these data it is apparent that the distal regions of some RNA polymerase III- and RNA polymerase II-transcribed small RNA promoters are virtually identical in composition, and their mechanisms of transcriptional activation are possibly quite similar.
Figures






References
-
- Ares M. Jr.; Chung J.-S.; Giglio L.; Weiner A. M. Distinct factors with Spl and NF-A specificities bind to adjacent functional elements of the human U2 snRNA gene enhancer. Genes Dev. 1:808–817; 1987. - PubMed
-
- Christensen J. H.; Hansen P. K.; Lillelund O.; Thogersen H. C. Sequence-specific binding of the N-terminal three-finger fragment of Xenopus transcription factor IIIA to the internal control region of a 5S RNA gene. FEBS Lett. 281:181–184; 1991. - PubMed
-
- Dahlberg J. E.; Lund E. How does III × II make U6. Science 254:1462–1463; 1991. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
