Capsule switching of Neisseria meningitidis
- PMID: 8990198
- PMCID: PMC19312
- DOI: 10.1073/pnas.94.1.271
Capsule switching of Neisseria meningitidis
Abstract
The different sialic acid (serogroups B, C, Y, and W-135) and nonsialic acid (serogroup A) capsular polysaccharides expressed by Neisseria meningitidis are major virulence factors and are used as epidemiologic markers and vaccine targets. However, the identification of meningococcal isolates with similar genetic markers but expressing different capsular polysaccharides suggests that meningococcal clones can switch the type of capsule they express. We identified, except for capsule, isogenic serogroups B [(alpha2-->8)-linked polysialic acid] and C [(alpha2-->9)-linked polysialic acid] meningococcal isolates from an outbreak of meningococcal disease in the U. S. Pacific Northwest. We used these isolates and prototype serogroup A, B, C, Y, and W-135 strains to define the capsular biosynthetic and transport operons of the major meningococcal serogroups and to show that switching from the B to C capsule in the outbreak strain was the result of allelic exchange of the polysialyltransferase. Capsule switching was probably the result of transformation and horizontal DNA exchange in vivo of a serogroup C capsule biosynthetic operon. These findings indicate that closely related virulent meningococcal clones may not be recognized by traditional serogroup-based surveillance and can escape vaccine-induced or natural protective immunity by capsule switching. Capsule switching may be an important virulence mechanism of meningococci and other encapsulated bacterial pathogens. As vaccine development progresses and broader immunization with capsular polysaccharide conjugate vaccines becomes a reality, the ability to switch capsular types may have important implications for the impact of these vaccines.
Figures
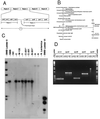





References
-
- Apicella M A. In: Principles and Practice of Infectious Diseases. Mandell G L, Douglas R G, Bennett J E, editors. New York: Churchill Livingstone; 1995. pp. 1896–1909.
-
- W.H.O. Fighting Disease, Fostering Development. Geneva: W.H.O.; 1996.
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

