The yeast Red1 protein localizes to the cores of meiotic chromosomes
- PMID: 9060462
- PMCID: PMC2132480
- DOI: 10.1083/jcb.136.5.957
The yeast Red1 protein localizes to the cores of meiotic chromosomes
Abstract
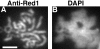
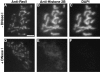
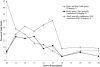
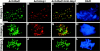
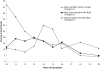
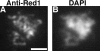
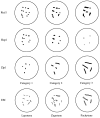
Mutants in the meiosis-specific RED1 gene of S. cerevisiae fail to make any synaptonemal complex (SC) or any obvious precursors to the SC. Using antibodies that specifically recognize the Red1 protein, Red1 has been localized along meiotic pachytene chromosomes. Red1 also localizes to the unsynapsed axial elements present in a zip1 mutant, suggesting that Red1 is a component of the lateral elements of mature SCs. Anti-Red1 staining is confined to the cores of meiotic chromosomes and is not associated with the loops of chromatin that lie outside the SC. Analysis of the spo11 mutant demonstrates that Red1 localization does not depend upon meiotic recombination. The localization of Red1 has been compared with two other meiosis-specific components of chromosomes, Hop1 and Zip1; Zip1 serves as a marker for synapsed chromosomes. Double labeling of wild-type meiotic chromosomes with anti-Zip1 and anti-Red1 antibodies demonstrates that Red1 localizes to chromosomes both before and during pachytene. Double labeling with anti-Hop1 and anti-Red1 antibodies reveals that Hop1 protein localizes only in areas that also contain Red1, and studies of Hop1 localization in a red1 null mutant demonstrate that Hop1 localization depends on Red1 function. These observations are consistent with previous genetic studies suggesting that Red1 and Hop1 directly interact. There is little or no Hop1 protein on pachytene chromosomes or in synapsed chromosomal regions.
Figures




References
-
- Alani E, Padmore R, Kleckner N. Analysis of wild-type and rad50mutants of yeast suggests an intimate relationship between meiotic chromosome synapsis and recombination. Cell. 1990;61:419–436. - PubMed
-
- Bishop DK. RecA homologues Dmc1 and Rad51 interact to form discrete nuclear complexes prior to meiotic chromosome synapsis. Cell. 1994;79:1081–1092. - PubMed
-
- Cao L, Alani E, Kleckner N. A pathway for generation and processing of double-strand breaks during meiotic recombination in S. cerevisiae. . Cell. 1990;61:1089–1101. - PubMed
-
- Dobson MJ, Pearlman RE, Karaiskakis A, Spyropoulos B, Moens PB. Synaptonemal complex proteins: occurrence, epitope mapping and chromosome disjunction. J Cell Sci. 1994;107:2749–2760. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

