Actin depolymerizing factor (ADF/cofilin) enhances the rate of filament turnover: implication in actin-based motility
- PMID: 9087445
- PMCID: PMC2132522
- DOI: 10.1083/jcb.136.6.1307
Actin depolymerizing factor (ADF/cofilin) enhances the rate of filament turnover: implication in actin-based motility
Abstract
Actin-binding proteins of the actin depolymerizing factor (ADF)/cofilin family are thought to control actin-based motile processes. ADF1 from Arabidopsis thaliana appears to be a good model that is functionally similar to other members of the family. The function of ADF in actin dynamics has been examined using a combination of physical-chemical methods and actin-based motility assays, under physiological ionic conditions and at pH 7.8. ADF binds the ADP-bound forms of G- or F-actin with an affinity two orders of magnitude higher than the ATP- or ADP-Pi-bound forms. A major property of ADF is its ability to enhance the in vitro turnover rate (treadmilling) of actin filaments to a value comparable to that observed in vivo in motile lamellipodia. ADF increases the rate of propulsion of Listeria monocytogenes in highly diluted, ADF-limited platelet extracts and shortens the actin tails. These effects are mediated by the participation of ADF in actin filament assembly, which results in a change in the kinetic parameters at the two ends of the actin filament. The kinetic effects of ADF are end specific and cannot be accounted for by filament severing. The main functionally relevant effect is a 25-fold increase in the rate of actin dissociation from the pointed ends, while the rate of dissociation from the barbed ends is unchanged. This large increase in the rate-limiting step of the monomer-polymer cycle at steady state is responsible for the increase in the rate of actin-based motile processes. In conclusion, the function of ADF is not to sequester G-actin. ADF uses ATP hydrolysis in actin assembly to enhance filament dynamics.
Figures







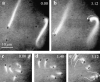



Comment in
-
Accelerating on a treadmill: ADF/cofilin promotes rapid actin filament turnover in the dynamic cytoskeleton.J Cell Biol. 1997 Mar 24;136(6):1165-8. doi: 10.1083/jcb.136.6.1165. J Cell Biol. 1997. PMID: 9087434 Free PMC article. Review. No abstract available.
References
-
- Abe H, Obinata T. An actin-depolymerizing protein in embryonic chicken skeletal muscle: purification and characterization. J Biochem. 1989;106:172–180. - PubMed
-
- Abe H, Oshima S, Obinata T. A cofilin-like protein is involved in the regulation of actin assembly in developing skeletal muscle. J Biochem. 1989;106:696–702. - PubMed
-
- Agnew BJ, Minamide LS, Bamburg JR. Reactivation of phosphorylated actin depolymerizing factor and identification of the regulatory site. J Biol Chem. 1995;270:17582–17587. - PubMed
-
- Aizawa H, Sutoh K, Tsubuki S, Kawashima S, Ishii A, Yahara I. Identification, characterization and intracellular distribution of cofilin in Dictyostelium discoideum. . J Biol Chem. 1995;270:10923–10932. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

