Quantitative trait loci involved in genetic predisposition to acute alcohol withdrawal in mice
- PMID: 9133412
- PMCID: PMC6573712
- DOI: 10.1523/JNEUROSCI.17-10-03946.1997
Quantitative trait loci involved in genetic predisposition to acute alcohol withdrawal in mice
Abstract
Alcohol dependence (alcoholism) is accompanied by evidence of tolerance, withdrawal (physiological dependence), or compulsive behavior related to alcohol use. Studies of strain and individual differences using animal models for acute physiological dependence liability are useful means to identify potential genetic determinants of liability in humans. Behavioral and quantitative trait analyses were conducted using animal models for high risk versus resistance to acute physiological dependence. Using a two-step genetic mapping strategy, loci on mouse chromosomes 1, 4, and 11 were mapped that contain genes that influence alcohol withdrawal severity. In the aggregate, these three risk markers accounted for 68% of the genetic variability in alcohol withdrawal. Candidate genes in proximity to the chromosome 11 locus include genes encoding the alpha1, alpha6, and gamma2 subunits of type-A receptors for the inhibitory neurotransmitter, GABA. In addition, suggestive linkage is indicated for two loci on mouse chromosome 2, one near Gad1 encoding glutamic acid decarboxylase, and the other near the El2 locus which influences the seizure phenotype in the neurological mutant strain El. The present analyses detect and map some of the loci that increase risk to develop physiological dependence and may facilitate identification of genes related to the development of alcoholism. Syntenic conservation between human and mouse chromosomes suggests that human homologs of genes that increase risk for physiological dependence may localize to 1q21-q32, 2q24-q37/11p13, 9p21-p23/1p32-p22.1, and 5q32-q35.
Figures

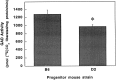
References
-
- Albers RW, Brady RO. The distribution of glutamic decarboxylase in the nervous system of the monkey. J Biol Chem. 1959;234:926–928. - PubMed
-
- Belknap JK, Laursen SE, Crabbe JC. Ethanol and nitrous oxide produce withdrawal-induced convulsions by similar mechanisms in mice. Life Sciences. 1987;41:2033–2040. - PubMed
-
- Belknap JK, Danielson PW, Lame M, Crabbe JC. Ethanol and barbiturate withdrawal convulsions are extensively codetermined in mice. Alcohol. 1988;5:167–171. - PubMed
-
- Belknap JK, Crabbe JC, Laursen SE. Ethanol and diazepam withdrawal convulsions are extensively codetermined in WSP and WSR mice. Life Sciences. 1989;44:2075–2080. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
