The herpes simplex virus 1 protein kinase US3 is required for protection from apoptosis induced by the virus
- PMID: 9223283
- PMCID: PMC21525
- DOI: 10.1073/pnas.94.15.7891
The herpes simplex virus 1 protein kinase US3 is required for protection from apoptosis induced by the virus
Abstract
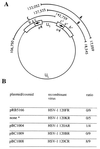
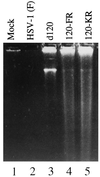
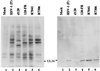
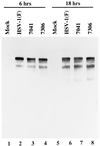
An earlier report showed that a disabled mutant lacking both copies of the major regulatory gene (alpha4) of herpes simplex virus 1 induced DNA degradation characteristic of apoptosis in infected cells, whereas the wild-type virus protected cells from apoptosis induced by thermal shock. More extensive analyses of the disabled mutant revealed a second mutation which disabled US3, a viral gene encoding a protein kinase known to phosphorylate serine/threonine within a specific arginine-rich consensus sequence. Analyses of cells infected with a viral mutant carrying a wild-type alpha4 gene but from which the US3 gene had been deleted showed that it induced fragmentation of cellular DNA, whereas a recombinant virus in which the deleted sequences of the US3 gene had been restored did not cause the cellular DNA to fragment. These results point to the protein kinase encoded by the US3 gene as the principal viral product required to block apoptosis.
Figures


References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

